Structural Dynamics of OATP1A2 in Mediating Paclitaxel Transport Mechanism in Breast Cancer
- PMID: 40078312
- PMCID: PMC11898720
- DOI: 10.7150/ntno.103095
Structural Dynamics of OATP1A2 in Mediating Paclitaxel Transport Mechanism in Breast Cancer
Abstract
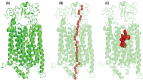
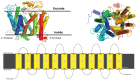
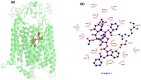
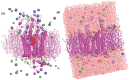
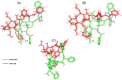
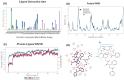
Breast cancer remains a significant global health challenge, with drug resistance and poor bioavailability of chemotherapeutic agents like paclitaxel (PTX) presenting obstacles to effective treatment. This study investigates the potential role of the Solute Carrier Organic Anion Transporter Polypeptide 1A2 (OATP1A2) in PTX transport using computational approaches. We employed computational modeling, molecular docking, and molecular dynamics (MD) simulations to elucidate the structural dynamics of OATP1A2 and its interaction with PTX. The OATP1A2 structure was modeled using Phyre2, validated, and refined. Molecular docking revealed significant PTX interactions within the predicted binding site, with a binding affinity of -10.4 kcal/mol and initial hydrogen bonding with Arg656 and Gly560 and hydrophobic interaction with atGlu66, Phe65, Asn41, Ala203, Ile204, Phe329, Phe332, Ile336, Pro207, Ser337, Asn334. Contrary to our initial hypothesis of inward drug movement, MD simulation over 500 ns revealed an unexpected outward movement of PTX. The ligand shifted approximately 5.4 Å towards the extracellular side from its initial binding position. This observation suggests a more complex transport mechanism than initially anticipated. The protein-ligand complex exhibited stability throughout the simulation, with notable conformational changes. Our findings highlight the complex nature of OATP1A2-mediated transport and its potential limitations for PTX delivery. These results accentuate the complexity of transporter-mediated drug delivery and may inform future strategies for improving chemotherapeutic efficacy in breast cancer treatment.
Keywords: Membrane Transporter; Molecular Dynamics; OATP1A2; Paclitaxel.; Protein Homology Modeling; Protein-ligand Docking.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures
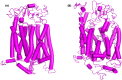
Similar articles
-
Solute Carrier Family of the Organic Anion-Transporting Polypeptides 1A2- Madin-Darby Canine Kidney II: A Promising In Vitro System to Understand the Role of Organic Anion-Transporting Polypeptide 1A2 in Blood-Brain Barrier Drug Penetration.Drug Metab Dispos. 2015 Jul;43(7):1008-18. doi: 10.1124/dmd.115.064170. Epub 2015 Apr 23. Drug Metab Dispos. 2015. PMID: 25908246
-
Influence of human OATP1B1, OATP1B3, and OATP1A2 on the pharmacokinetics of methotrexate and paclitaxel in humanized transgenic mice.Clin Cancer Res. 2013 Feb 15;19(4):821-32. doi: 10.1158/1078-0432.CCR-12-2080. Epub 2012 Dec 14. Clin Cancer Res. 2013. PMID: 23243220
-
Neurosteroids: Structure-Uptake Relationships and Computational Modeling of Organic Anion Transporting Polypeptides (OATP)1A2.Molecules. 2021 Sep 17;26(18):5662. doi: 10.3390/molecules26185662. Molecules. 2021. PMID: 34577133 Free PMC article.
-
pH-dependent transport kinetics of the human organic anion-transporting polypeptide 1A2.Drug Metab Pharmacokinet. 2020 Apr;35(2):220-227. doi: 10.1016/j.dmpk.2019.12.002. Epub 2019 Dec 24. Drug Metab Pharmacokinet. 2020. PMID: 31974044
-
Genetic polymorphisms and function of the organic anion-transporting polypeptide 1A2 and its clinical relevance in drug disposition.Pharmacology. 2015;95(3-4):201-8. doi: 10.1159/000381313. Epub 2015 Apr 23. Pharmacology. 2015. PMID: 25924632 Review.
References
-
- Iacoviello L, Bonaccio M, de Gaetano G, Donati MB. Epidemiology of breast cancer, a paradigm of the “common soil” hypothesis: Seminars Semin Cancer Biol. 2021;72:4-10. - PubMed
-
- Zhang B, Lauschke VM. Genetic variability and population diversity of the human SLCO (OATP) transporter family. Pharmaco Res. 2019;139:550–9. - PubMed
-
- Svoboda M, Riha J, Wlcek K, Jaeger W, Thalhammer T. Organic anion transporting polypeptides (OATPs): regulation of expression and function. Curr Drug Metab. 2011;12(2):139–53. - PubMed
-
- Hagenbuch B, Meier PJ. The superfamily of organic anion transporting polypeptides. BiochimBiophys Acta-Biomem. 2003;1609(1):1–18. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

