AFM reveals differential effects of acidification on LDL- and oxidized LDL-receptor interactions: biomechanical implications in atherogenesis
- PMID: 40102716
- PMCID: PMC11921551
- DOI: 10.1186/s11658-025-00715-9
AFM reveals differential effects of acidification on LDL- and oxidized LDL-receptor interactions: biomechanical implications in atherogenesis
Abstract
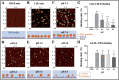
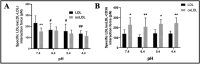
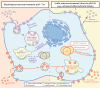
The receptor recognition and interaction of plasma lipoproteins (e.g., native low-density lipoproteins (LDL)/oxidized low-density lipoproteins (oxLDL), as well as the influence of microenvironmental/lysosomal acidification, play critical roles in lipoprotein metabolism and diseases (e.g., atherosclerosis) but have been less investigated. Here, the recognition/interaction of LDL or oxLDL with LDL receptor (LDLR) or CD36 (a scavenger receptor) or with living cells at various pHs was evaluated mainly via atomic force microscopy (AFM). To improve force measurement accuracy, a novel, micro-droplet-based method for AFM probe functionalization was developed. We found that solution acidification significantly reduced the LDL-LDLR binding at pH ≤ 6.4, whereas the oxLDL-CD36 binding had no significant change until pH ≤ 4.4. Compared with a traditional immersion method, our micro-droplet method for AFM probe functionalization produced more accurate interaction forces, and revealed that acidification significantly reduced the LDL-LDLR/cell interaction forces, instead of the oxLDL-CD36/cell-specific interaction forces and nonspecific interaction forces. The data imply that the LDL-LDLR/cell recognition and interaction are susceptible to acidification, whereas the oxLDL-CD36/cell recognition and interaction are tolerant of acidification. The results may provide important novel information and biomechanical/pathological implications for understanding lipoprotein metabolism and atherosclerosis.
Keywords: Atherosclerosis; Atomic force microscopy (AFM); Low-density lipoprotein (LDL).
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: All authors approved the publication. Competing interests: The authors declare no competing interests.
Figures






Similar articles
-
AFM detects the effects of acidic condition on the size and biomechanical properties of native/oxidized low-density lipoprotein.Colloids Surf B Biointerfaces. 2021 Dec;208:112053. doi: 10.1016/j.colsurfb.2021.112053. Epub 2021 Aug 19. Colloids Surf B Biointerfaces. 2021. PMID: 34438294
-
Native and modified low density lipoproteins increase the functional expression of the macrophage class B scavenger receptor, CD36.J Biol Chem. 1997 Aug 22;272(34):21654-9. doi: 10.1074/jbc.272.34.21654. J Biol Chem. 1997. PMID: 9261189
-
Comparative investigation on the sizes and scavenger receptor binding of human native and modified lipoprotein particles with atomic force microscopy.J Nanobiotechnology. 2018 Mar 21;16(1):25. doi: 10.1186/s12951-018-0352-3. J Nanobiotechnology. 2018. PMID: 29592798 Free PMC article.
-
Foam cells in atherosclerosis.Clin Chim Acta. 2013 Sep 23;424:245-52. doi: 10.1016/j.cca.2013.06.006. Epub 2013 Jun 16. Clin Chim Acta. 2013. PMID: 23782937 Review.
-
The enigmatic membrane fatty acid transporter CD36: New insights into fatty acid binding and their effects on uptake of oxidized LDL.Prostaglandins Leukot Essent Fatty Acids. 2018 Nov;138:64-70. doi: 10.1016/j.plefa.2016.05.005. Epub 2016 May 20. Prostaglandins Leukot Essent Fatty Acids. 2018. PMID: 27288302 Review.
References
-
- Barth JD, Arntzenius AC. Progression and regression of atherosclerosis, what roles for LDL-cholesterol and HDL-cholesterol—a perspective. Eur Heart J. 1991;12(8):952–7. - PubMed
-
- Qin SC. LDL and HDL Oxidative modification and atherosclerosis. In: Jiang XC, editor. Lipid transfer in lipoprotein metabolism and cardiovascular disease, vol. 1276. Singapore: Springer; 2020. p. 157–69. - PubMed
-
- Ao MY, Gan CY, Shao WX, Zhou X, Chen Y. Effects of cyclodextrins on the structure of LDL and its susceptibility to copper-induced oxidation. Eur J Pharm Sci. 2016;91:183–9. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

