Enable, empower, succeed: a bioinformatics workshop Harnessing open web-based tools for surveillance of bacterial antimicrobial resistance
- PMID: 40102762
- PMCID: PMC11921729
- DOI: 10.1186/s12866-025-03865-0
Enable, empower, succeed: a bioinformatics workshop Harnessing open web-based tools for surveillance of bacterial antimicrobial resistance
Abstract
Background: Antimicrobial resistance (AMR) poses a significant threat to global health, particularly in Western sub-Saharan Africa where 27.3 deaths per 100,000 lives are affected, and surveillance and control measures are often limited. Genomics research plays a crucial role in understanding the emergence, spread and containment measures of AMR. However, its implementation in such settings is particularly challenging due to limited human capacity. This manuscript outlines a three-day bioinformatics workshop in Cameroon, highlighting efforts to build human capacity for genomics research to support AMR surveillance using readily accessible and user-friendly web-based tools. The workshop introduced participants to basic next-generation sequencing concepts, data file formats used in bacterial genomics, data sharing procedures and considerations, as well as the use of web-based bioinformatics software to analyse genomic data, including in silico prediction of AMR, phylogenetics analyses, and a quick introduction to Linux© command line.
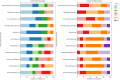
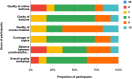
Results: Briefly, a substantial increase in participants' confidence in bioinformatics knowledge and skills was observed before and after the workshop. Notably, before the workshop most participants lacked confidence in their ability to identify next-generation sequencing technologies or workflows (64%) and analyse genetic data using web-based bioinformatics tools (81%). After the workshop, majority of participants were extremely confident using NCBI BLAST and other web-based bioinformatics tools for data analysis with a score ≥ 5 among which 45%, 9% and 18% had a score of 8, 9, and 10, respectively.
Conclusion: Our findings highlight the effectiveness of this training approach in empowering local researchers and bridging the bioinformatics gap in genomics surveillance of AMR in resource-constrained settings. We provide a detailed description of the relevant training approaches used, including workshop structure, the selection and planning, and utilization of freely available web-based tools, and the evaluation methods employed. Our approach aimed to overcome limitations such as inadequate infrastructure, limited access to computational resources, and scarcity of expertise. By leveraging the power of freely available web-based tools, we demonstrated how participants can acquire fundamental bioinformatics skills, enhance their understanding of biological data analysis, and contribute to the field, even in an underprivileged environment. Building human capacity for genomics research globally, and especially in resource-constrained settings, is imperative for ensuring global health and sustainable containment of AMR.
Keywords: Africa; Antimicrobial resistance; Bioinformatics; Cameroon; Capacity Building; Resource-constrained settings; Skills development.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Human ethics and consent to participate: Institutional approval was obtained from the CEDBCAM-RI to utilize the anonymized data collected during the workshop. This study was conducted in accordance with the ethical principles outlined in the Declaration of Helsinki and approved by the National Ethics Committee for Human Health Research (No. 2021/07/1386/CE/CNERSH/SP) and the Ethical Committee of the University of Douala (No. 3190 CEI-UDo/06/2022/M). All participants provided written informed consent to have their data analyzed and published as part of this study. Consent for publication: Not applicable. Conflict of interest: The authors declare that there are no conflicts of interest. Clinical trial number: Not applicable. Impact statement: Antimicrobial resistance (AMR) is a major global health threat, especially in Western sub-Saharan Africa, where 27.3 deaths per 100,000 lives occur. Genomics research play an instrumental role for understanding AMR's emergence, spread, and containment measures. However, its implementation in these settings is challenging due to limited human capacity. A three-day bioinformatics workshop in Cameroon aimed to build human capacity for genomics research using web-based tools. Participants were introduced to next-generation sequencing concepts, data file formats, data sharing procedures, and web-based bioinformatics software for analysing genomic data. The workshop aimed to overcome limitations like inadequate infrastructure, computational resources, and expertise scarcity. The findings show the effectiveness of this training approach in empowering local researchers and bridging the bioinformatics gap in genomics surveillance of AMR in resource-constrained settings.
Figures
Similar articles
-
AMRColab - a user-friendly antimicrobial resistance detection and visualization tool.Microb Genom. 2024 Oct;10(10):001308. doi: 10.1099/mgen.0.001308. Microb Genom. 2024. PMID: 39432417 Free PMC article.
-
Assessment of the pathogen genomics landscape highlights disparities and challenges for effective AMR Surveillance and outbreak response in the East African community.BMC Public Health. 2024 Jun 5;24(1):1500. doi: 10.1186/s12889-024-18990-0. BMC Public Health. 2024. PMID: 38840103 Free PMC article.
-
Analysis of Whole Genome Sequencing Data for Detection of Antimicrobial Resistance Determinants.Methods Mol Biol. 2024;2833:211-223. doi: 10.1007/978-1-0716-3981-8_19. Methods Mol Biol. 2024. PMID: 38949713
-
Genomics for antimicrobial resistance surveillance to support infection prevention and control in health-care facilities.Lancet Microbe. 2023 Dec;4(12):e1040-e1046. doi: 10.1016/S2666-5247(23)00282-3. Epub 2023 Nov 14. Lancet Microbe. 2023. PMID: 37977161 Review.
-
Antimicrobial resistance surveillance in the genomic age.Ann N Y Acad Sci. 2017 Jan;1388(1):78-91. doi: 10.1111/nyas.13289. Epub 2016 Nov 22. Ann N Y Acad Sci. 2017. PMID: 27875856 Review.
References
-
- O’Neill J. Tackling Drug-resistant infections globally: final report and recommendations. Review on Antimicrobial Resistance; 2016.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous