Modular access to nucleobase GFP-surrogates: pH-responsive smart probes for ratiometric nucleic acid diagnostics
- PMID: 40103717
- PMCID: PMC11912499
- DOI: 10.1039/d4sc07994a
Modular access to nucleobase GFP-surrogates: pH-responsive smart probes for ratiometric nucleic acid diagnostics
Abstract
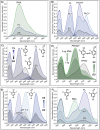
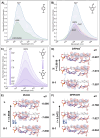
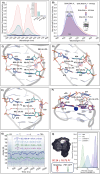
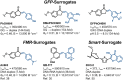
We have utilized a modular on-strand aldol approach to synthesize chalcone-based fluorescent molecular rotors (FMRs) bearing phenolic oxygen donors that mimic the natural tyrosine (Tyr66) chromophore 4-hydroxybenzylidene-imidazolinone (HBI) within green fluorescent proteins (GFPs). Leveraging the FMRs' propensity to undergo non-radiative decay via twisted intramolecular charge transfer upon excitation within certain microenvironments, we have addressed the longstanding issues of poor brightness (ε max × Φ fl) and weak turn-on responses for GFP-surrogates within nucleic acids. To demonstrate its potential and lay the groundwork for future applications, these FMRs were incorporated into NarI12 and TBA15 oligonucleotides with canonical (A, C, T, G) or locked nucleic acids (LNAs) (TL, AL) as flanking bases. The resulting duplexes and G-quadruplexes (GQs) were studied using fluorescence spectroscopy, molecular dynamics simulations, and quantum mechanical calculations, yielding a comprehensive understanding of their structural and photophysical properties in DNA, DNA : RNA, and GQ contexts. Electron-rich chalcones favor neutral phenol excitation (ROH) to afford both phenol (ROH*) and phenolate (RO-*) emission, with the latter generated through an intermolecular excited-state proton transfer process, while electron-deficient chalcones serve as ratiometric excitation indicators, due to their photoacidity. The surrogates display strong turn-on responses (up to 154-fold) in a GQ → duplex topology switch with flanked LNAs, giving Φ fl up to 0.58 and molar brightness ∼ 15 000 cm-1 M-1 in the duplex. By synergizing the NA sequence and probe, we achieve a switchable ON-to-OFF photoinduced electron transfer, resulting in a 134-fold turn-on emission response to pH. Our findings are the first to optimize the performance of GFP-surrogates as internal nucleobase replacements and suggest multiple ways in which they may be useful tools for NA diagnostics.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
There are no conflicts to declare.
Figures



Similar articles
-
A modular aldol approach for internal fluorescent molecular rotor chalcone surrogates for DNA biosensing applications.Chem Sci. 2023 Apr 11;14(18):4832-4844. doi: 10.1039/d3sc00772c. eCollection 2023 May 10. Chem Sci. 2023. PMID: 37181758 Free PMC article.
-
Harnessing a 4-Formyl-Aniline Handle to Tune the Stability of a DNA Aptamer-Protein Complex via Fluorescent Surrogates.Bioconjug Chem. 2023 Nov 15;34(11):2066-2076. doi: 10.1021/acs.bioconjchem.3c00373. Epub 2023 Oct 19. Bioconjug Chem. 2023. PMID: 37857354
-
5'-Amino-Formyl-Thieno[3,2-b]thiophene End-Label for On-Strand Synthesis of Far-Red Fluorescent Molecular Rotors and pH-Responsive Probes.Bioconjug Chem. 2025 Feb 19;36(2):216-222. doi: 10.1021/acs.bioconjchem.4c00457. Epub 2025 Jan 21. Bioconjug Chem. 2025. PMID: 39838819
-
Fundamentals of the Intrinsic DNA Fluorescence.Acc Chem Res. 2021 Mar 2;54(5):1226-1235. doi: 10.1021/acs.accounts.0c00603. Epub 2021 Feb 15. Acc Chem Res. 2021. PMID: 33587613 Review.
-
Pyrene-functionalized oligonucleotides and locked nucleic acids (LNAs): tools for fundamental research, diagnostics, and nanotechnology.Chem Soc Rev. 2011 Dec;40(12):5771-88. doi: 10.1039/c1cs15014f. Epub 2011 Apr 13. Chem Soc Rev. 2011. PMID: 21487621 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

