AlgaeOrtho, a bioinformatics tool for processing ortholog inference results in algae
- PMID: 40104598
- PMCID: PMC11913701
- DOI: 10.3389/fmicb.2025.1541898
AlgaeOrtho, a bioinformatics tool for processing ortholog inference results in algae
Abstract
Introduction: Microalgae constitute a prominent feedstock for producing biofuels and biochemicals by virtue of their prolific reproduction, high bioproduct accumulation, and the ability to grow in brackish and saline water. However, naturally occurring wild type algal strains are rarely optimal for industrial use; therefore, bioengineering of algae is necessary to generate superior performing strains that can address production challenges in industrial settings, particularly the bioenergy and bioproduct sectors. One of the crucial steps in this process is deciding on a bioengineering target: namely, which gene/protein to differentially express. These targets are often orthologs which are defined as genes/proteins originating from a common ancestor in divergent species. Although bioinformatics tools for the identification of protein orthologs already exist, processing the output from such tools is nontrivial, especially for a researcher with little or no bioinformatics experience.
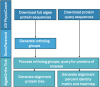
Methods: The present study introduces AlgaeOrtho, a user-friendly tool that builds upon the SonicParanoid orthology inference tool (based on an algorithm that identifies potential protein orthologs based on amino acid sequences) and the PhycoCosm database from JGI (Joint Genome Institute) to help researchers identify orthologs of their proteins of interest in multiple diverse algal species.
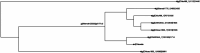
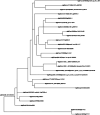
Results: The output of this application includes a table of the putative orthologs of their protein of interest, a heatmap showing sequence similarity (%), and an unrooted tree of the putative protein orthologs. Notably, the tool would be instrumental in identifying novel bioengineering targets in different algal strains, including targets in not-fully annotated algal species, since it does not depend on existing protein annotations. We tested AlgaeOrtho using three case studies, for which orthologs of proteins relevant to bioengineering targets, were identified from diverse algal species, demonstrating its ease of use and utility for bioengineering researchers.
Discussion: This tool is unique in the protein ortholog identification space as it can visualize putative orthologs, as desired by the user, across several algal species.
Keywords: algae; bioengineering; bioinformatics; metabolic engineering; nutraceuticals; protein orthology.
Copyright © 2025 LaPorte, Arora, Clark and Nag.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
KinOrtho: a method for mapping human kinase orthologs across the tree of life and illuminating understudied kinases.BMC Bioinformatics. 2021 Sep 18;22(1):446. doi: 10.1186/s12859-021-04358-3. BMC Bioinformatics. 2021. PMID: 34537014 Free PMC article.
-
Automatic clustering of orthologs and in-paralogs from pairwise species comparisons.J Mol Biol. 2001 Dec 14;314(5):1041-52. doi: 10.1006/jmbi.2000.5197. J Mol Biol. 2001. PMID: 11743721
-
PhycoCosm, a comparative algal genomics resource.Nucleic Acids Res. 2021 Jan 8;49(D1):D1004-D1011. doi: 10.1093/nar/gkaa898. Nucleic Acids Res. 2021. PMID: 33104790 Free PMC article.
-
Bioengineering strategies of microalgae biomass for biofuel production: recent advancement and insight.Bioengineered. 2023 Dec;14(1):2252228. doi: 10.1080/21655979.2023.2252228. Bioengineered. 2023. PMID: 37661811 Free PMC article. Review.
-
Harnessing algal biomass for sustainable energy: cultivation, strain improvement, and biofuel production.Prep Biochem Biotechnol. 2025;55(5):521-534. doi: 10.1080/10826068.2024.2434879. Epub 2024 Dec 16. Prep Biochem Biotechnol. 2025. PMID: 39679595 Review.
References
-
- Araújo R., Vázquez Calderón F., Sánchez López J., Azevedo I. C., Bruhn A., Fluch S., et al. . (2021). Current status of the algae production industry in Europe: an emerging sector of the blue bioeconomy. Front. Mar. Sci. 7:626389. doi: 10.3389/fmars.2020.626389 - DOI
LinkOut - more resources
Full Text Sources

