Structure and dynamics determine G protein coupling specificity at a class A GPCR
- PMID: 40106559
- PMCID: PMC12101717
- DOI: 10.1126/sciadv.adq3971
Structure and dynamics determine G protein coupling specificity at a class A GPCR
Erratum in
-
Erratum for the Research Article "Structure and dynamics determine G protein coupling specificity at a class A GPCR" by M. Casiraghi et al.Sci Adv. 2025 Jun 27;11(26):eadz4217. doi: 10.1126/sciadv.adz4217. Epub 2025 Jun 27. Sci Adv. 2025. PMID: 40577483 Free PMC article. No abstract available.
Abstract
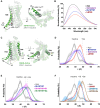
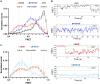
G protein-coupled receptors (GPCRs) exhibit varying degrees of selectivity for different G protein isoforms. Despite the abundant structures of GPCR-G protein complexes, little is known about the mechanism of G protein coupling specificity. The β2-adrenergic receptor is an example of GPCR with high selectivity for Gαs, the stimulatory G protein for adenylyl cyclase, and much weaker for the Gαi family of G proteins inhibiting adenylyl cyclase. By developing a Gαi-biased agonist (LM189), we provide structural and biophysical evidence supporting that distinct conformations at ICL2 and TM6 are required for coupling of the different G protein subtypes Gαs and Gαi. These results deepen our understanding of G protein specificity and bias and can accelerate the design of ligands that select for preferred signaling pathways.
Figures




Update of
-
Structure and dynamics determine G protein coupling specificity at a class A GPCR.bioRxiv [Preprint]. 2025 Jan 31:2024.03.28.587240. doi: 10.1101/2024.03.28.587240. bioRxiv. 2025. Update in: Sci Adv. 2025 Mar 21;11(12):eadq3971. doi: 10.1126/sciadv.adq3971. PMID: 38586060 Free PMC article. Updated. Preprint.
References
-
- Fredriksson R., Lagerström M. C., Lundin L.-G., Schiöth H. B., The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol. Pharmacol. 63, 1256–1272 (2003). - PubMed
-
- Koehl A., Hu H., Maeda S., Zhang Y., Qu Q., Paggi J. M., Latorraca N. R., Hilger D., Dawson R., Matile H., Schertler G. F. X., Granier S., Weis W. I., Dror R. O., Manglik A., Skiniotis G., Kobilka B. K., Structure of the μ-opioid receptor–Gi protein complex. Nature 558, 547–552 (2018). - PMC - PubMed
-
- Rasmussen S. G. F., DeVree B. T., Zou Y., Kruse A. C., Chung K. Y., Kobilka T. S., Thian F. S., Chae P. S., Pardon E., Calinski D., Mathiesen J. M., Shah S. T. A., Lyons J. A., Caffrey M., Gellman S. H., Steyaert J., Skiniotis G., Weis W. I., Sunahara R. K., Kobilka B. K., Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011). - PMC - PubMed
-
- Krishna Kumar K., Shalev-Benami M., Robertson M. J., Hu H., Banister S. D., Hollingsworth S. A., Latorraca N. R., Kato H. E., Hilger D., Maeda S., Weis W. I., Farrens D. L., Dror R. O., Malhotra S. V., Kobilka B. K., Skiniotis G., Structure of a signaling cannabinoid receptor 1-G protein complex. Cell 176, 448–458.e12 (2019). - PMC - PubMed

