The ETHYLENE RESPONSE FACTOR6-GRETCHEN HAGEN3.5 module regulates rooting and heat tolerance in Dimocarpus longan
- PMID: 40106655
- PMCID: PMC11950727
- DOI: 10.1093/plphys/kiaf096
The ETHYLENE RESPONSE FACTOR6-GRETCHEN HAGEN3.5 module regulates rooting and heat tolerance in Dimocarpus longan
Abstract
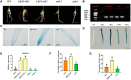
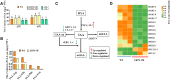
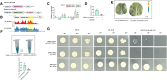
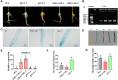
Heat stress can seriously affect plant growth and development. Ethylene response factors (ERFs) play important roles in plant development and physiological responses. Here, we identified DlERF6, an ERF family transcription factor that promotes heat tolerance in Dimocarpus longan. DlERF6 was strongly induced by heat stress and IAA treatment in longan roots. Overexpression of DlERF6 generated abundant, fast-growing hairy roots and enhanced longan heat stress tolerance by promoting IAA biosynthesis and reactive oxygen species (ROS) scavenging. Additional assays indicated that DlERF6 directly binds to the DlGH3.5 promoter and represses its expression. Overexpressing DlGH3.5 reduced hairy root number, root length, and heat tolerance, concomitant with a reduction in IAA content and ROS scavenging. Collectively, these results reveal the molecular mechanism through which the DlERF6-DlGH3.5 module regulates root growth and heat stress tolerance, providing a gene network that can be used for the genetic improvement of longan.
© The Author(s) 2025. Published by Oxford University Press on behalf of American Society of Plant Biologists.
Conflict of interest statement
Conflict of interest statement. None declared.
Figures


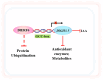
References
-
- Abiri R, Shaharuddin NA, Maziah M, Yusof ZNB, Atabaki N, Sahebi M, Valdiani A, Kalhori N, Azizi P, Hanafi MM. Role of ethylene and the APETALA 2/ethylene response factor superfamily in rice under various abiotic and biotic stress conditions. Environ Exp Bot. 2017:134:33–44. 10.1016/j.envexpbot.2016.10.015 - DOI
-
- Almeselmani M, Deshmukh PS, Sairam RK. High temperature stress tolerance in wheat genotypes: role of antioxidant defence enzymes. Acta Agron Hung. 2009:57(1):1–14. 10.1556/AAgr.57.2009.1.1 - DOI
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

