Revision of the Formycin A and Pyrazofurin Biosynthetic Pathways Reveals Specificity for d-Glutamic Acid and a Cryptic N-Acylation Step During Pyrazole Core Formation
- PMID: 40106796
- PMCID: PMC11964824
- DOI: 10.1021/jacs.5c01277
Revision of the Formycin A and Pyrazofurin Biosynthetic Pathways Reveals Specificity for d-Glutamic Acid and a Cryptic N-Acylation Step During Pyrazole Core Formation
Abstract
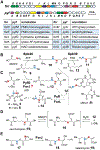
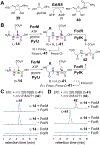
Formycin A and pyrazofurin are two naturally occurring pyrazole-derived C-nucleosides with antibacterial and antiviral activities. While earlier studies have established the chemistry of C-glycosidic bond formation as well as the subsequent steps in the biosynthesis of formycin A and pyrazofurin, how the pyrazole ring itself is constructed remains elusive. While N-N bond formation in the pyrazole ring was previously reported to involve coupling of N6-hydroxylated l-lysine and l-glutamic acid catalyzed by the hydrazine synthetase PyfG, herein PyfG and its homologue ForJ are shown instead to recognize d-glutamate instead of l-glutamate. The hydrazine product of ForJ/PyfG catalysis then releases α-hydrazino d-glutamic acid upon processing by the NAD-dependent oxidoreductase ForL. Furthermore, N-acylation of α-hydrazino d-glutamate with an amino acid catalyzed by the ATP-grasp ligase ForM/PyfJ is indispensable for recognition by the FAD-dependent oxidoreductase ForR/PyfK to perform dehydrogenation of the Cα-N bond and thereby form a hydrazone intermediate. This work not only demonstrates that d-glutamic acid is the correct substrate for hydrazine biosynthesis but also reveals a cryptic N-acylation step in the assembly of the pyrazole core. These results thus provide significant insights into the biosynthesis of pyrazole rings that are rarely seen in natural products.
Conflict of interest statement
The authors declare no competing financial interest.
Figures





Similar articles
-
Identification of the C-Glycoside Synthases during Biosynthesis of the Pyrazole-C-Nucleosides Formycin and Pyrazofurin.Angew Chem Int Ed Engl. 2019 Nov 11;58(46):16512-16516. doi: 10.1002/anie.201910356. Epub 2019 Oct 7. Angew Chem Int Ed Engl. 2019. PMID: 31518483 Free PMC article.
-
Biosynthesis of formycin. Incorporation and distribution of labeled compounds into formycin.J Antibiot (Tokyo). 1976 Jun;29(6):638-45. doi: 10.7164/antibiotics.29.638. J Antibiot (Tokyo). 1976. PMID: 950318
-
Identification of the Formycin A Biosynthetic Gene Cluster from Streptomyces kaniharaensis Illustrates the Interplay between Biological Pyrazolopyrimidine Formation and de Novo Purine Biosynthesis.J Am Chem Soc. 2019 Apr 17;141(15):6127-6131. doi: 10.1021/jacs.9b00241. Epub 2019 Apr 8. J Am Chem Soc. 2019. PMID: 30942582 Free PMC article.
-
New trends in synthesis of pyrazole nucleosides as new antimetabolites.Nucleosides Nucleotides Nucleic Acids. 2005;24(8):1227-47. doi: 10.1081/NCN-200067421. Nucleosides Nucleotides Nucleic Acids. 2005. PMID: 16270665 Review.
-
Recent Research Progress in Taxol Biosynthetic Pathway and Acylation Reactions Mediated by Taxus Acyltransferases.Molecules. 2021 May 12;26(10):2855. doi: 10.3390/molecules26102855. Molecules. 2021. PMID: 34065782 Free PMC article. Review.
Cited by
-
Identification of a l-Threonine-Utilizing Hydrazine Synthetase for Thrazarine Biosynthesis in Streptomyces coerulescens MH802-fF5.Chembiochem. 2025 Aug 22;26(15):e202500298. doi: 10.1002/cbic.202500298. Epub 2025 Jul 14. Chembiochem. 2025. PMID: 40568798 Free PMC article.
References
-
- Suhadolnik RJ Nucleoside Antibiotics; Wiley-Interscience, 1970.
-
- Gutowski GE; Sweeney MJ; DeLong DC; Hamill RL; Gerzon K; Dyke RW Biochemistry and biological effects of the pyrazofurins (pyrazomycins): initial clinical trial. Ann. N.Y. Acad. Sci 1975, 255, 544–551. - PubMed
-
- Hori M; Takita T; Koyama G; Tadeuchi T; Umezawa H A new antibiotic, formycin. J. Antibiot 1964, 17, 96–99. - PubMed
-
- Ochi K; Iwamoto S; Hayase E; Yashima S; Okami Y Biosynthesis of formycin. Role of certain amino acids in formycin biosynthesis. J. Antibiot 1974, 27 (12), 909–916. - PubMed
-
- Carson DA; Chang KP Phosphorylation and antileishmanial activity of formycin B. Biochem. Biophys. Res. Commun 1981, 100 (3), 1377–1383. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

