The lung microvasculature promotes alveolar type 2 cell differentiation via secreted SPARCL1
- PMID: 40118055
- PMCID: PMC12069885
- DOI: 10.1016/j.stemcr.2025.102451
The lung microvasculature promotes alveolar type 2 cell differentiation via secreted SPARCL1
Abstract
Lung endothelial cells (ECs) and pericytes are closely juxtaposed with the respiratory epithelium before birth and thus may have instructive roles during development. To test this hypothesis, we screened EC-secreted proteins for their ability to alter cell differentiation in alveolar organoids. We identified secreted protein acidic and rich in cysteine-like protein 1 (SPARCL1) as an extracellular matrix molecule that can promote alveolar type 2 (AT2) cell differentiation in vitro. SPARCL1-treated organoids display lysozyme upregulation and a doubling in the number of AT2 cells at the expense of intermediate progenitors. SPARCL1 also induces the upregulation of nuclear factor κB (NF-κB) target genes, and suppression of NF-κB activation in lung organoids blocked SPARCL1 effects. NF-κB activation by lipopolysaccharide (LPS) was sufficient to induce AT2 cell differentiation; however, pharmacological inhibition of the pathway alone did not prevent it. These data support a role for SPARCL1 and NF-κB in alveolar cell differentiation and suggest a potential value in targeting this signaling axis to promote alveolar maturation and regeneration.
Keywords: SPARCL1; alveolar type 2 cells; cell differentiation; endothelial cells; extracellular matrix; lung alveologenesis; lung development; lung organoids; pericytes; vascular-epithelial crosstalk.
Copyright © 2025 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures
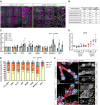



References
-
- Beauchemin K.J., Wells J.M., Kho A.T., Philip V.M., Kamir D., Kohane I.S., Graber J.H., Bult C.J. Temporal dynamics of the developing lung transcriptome in three common inbred strains of laboratory mice reveals multiple stages of postnatal alveolar development. PeerJ. 2016;4 doi: 10.7717/peerj.2318. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

