Non-canonical splice variants in thoracic aortic dissection cases and Marfan syndrome with negative genetic testing
- PMID: 40118890
- PMCID: PMC11928670
- DOI: 10.1038/s41525-025-00472-w
Non-canonical splice variants in thoracic aortic dissection cases and Marfan syndrome with negative genetic testing
Abstract
Individuals with heritable thoracic aortic disease (HTAD) face a high risk of deadly aortic dissections, but genetic testing identifies causative variants in only a minority of cases. We explored the contribution of non-canonical splice variants (NCVAS) to thoracic aortic disease (TAD) using SpliceAI and sequencing data from diverse cohorts, including 551 early-onset sporadic dissection cases and 437 HTAD probands with exome sequencing, 57 HTAD pedigrees with whole genome sequencing, and select sporadic cases with clinical panel testing. NCVAS were identified in syndromic HTAD genes such as FBN1, SMAD3, and COL3A1, including intronic variants in FBN1 in two Marfan syndrome (MFS) families. Validation in the Penn Medicine BioBank and UK Biobank showed enrichment of NCVAS in HTAD-associated genes among dissections. These findings suggest NCVAS are an underrecognized contributor to TAD, particularly in sporadic dissection and unsolved MFS cases, highlighting the potential of advanced splice prediction tools in genetic diagnostics.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures

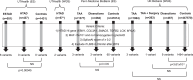
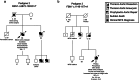
References
-
- Dietz, H. C. et al. Marfan syndrome caused by a recurrent de novo missense mutation in the fibrillin gene. Nature352, 337–339 (1991). - PubMed
-
- Silverman, D. I. et al. Life expectancy in the Marfan syndrome. Am. J. Cardiol.75, 157–160 (1995). - PubMed
-
- Harris, S. L. & Lindsay, M. E. Role of Clinical Genetic Testing in the Management of Aortopathies. Curr. Cardiol. Rep.23, 10 (2021). - PubMed
Grants and funding
- UL1 TR001878/TR/NCATS NIH HHS/United States
- U24 HG011746/HG/NHGRI NIH HHS/United States
- R01HL109942/U.S. Department of Health & Human Services | NIH | National Heart, Lung, and Blood Institute (NHLBI)
- 23POST1011251/American Heart Association (American Heart Association, Inc.)
- U01 HG011744/HG/NHGRI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Miscellaneous

