Design of orthogonal constant domain interfaces to aid proper heavy/light chain pairing of bispecific antibodies
- PMID: 40126074
- PMCID: PMC11934185
- DOI: 10.1080/19420862.2025.2479531
Design of orthogonal constant domain interfaces to aid proper heavy/light chain pairing of bispecific antibodies
Abstract
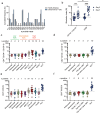
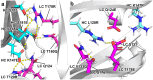
The correct pairing of cognate heavy and light chains is critical to the efficient manufacturing of IgG-like bispecific antibodies (bsAbs) from a single host cell. We present a general solution for the elimination of heavy chain (HC):light chain (LC) mispairs in bsAbs with LCs via the use of two orthogonal constant domain (CH1:C) interfaces comprising computationally designed amino acid substitutions. Substitutions were designed by Rosetta to introduce novel hydrogen bond (H-bond) networks at the CH1:C interface, followed by Rosetta energy calculations to identify designs with enhanced pairing specificity and interface stability. Our final design, featuring a total of 11 amino acid substitutions across two Fab constant regions, was tested on a set of six IgG-like bsAbs featuring a diverse set of unmodified human antibody variable domains. Purity assessments showed near-complete elimination of LC mispairs, including in cases with high baseline mispairing with wild-type constant domains. The engineered bsAbs broadly recapitulated the antigen-binding and biophysical developability properties of their monospecific counterparts and no adverse immunogenicity signal was identified by an in vitro assay. Fab crystal structures containing engineered constant domain interfaces revealed no major perturbations relative to the wild-type coordinates and validated the presence of the designed hydrogen bond interactions. Our work enables the facile assembly of independently discovered IgG-like bispecific antibodies in a single-cell host and demonstrates a streamlined and generalizable computational and experimental workflow for redesigning conserved protein:protein interfaces.
Keywords: Bispecific antibodies; antibody engineering; computational protein design; developability; heavy chain/light chain pairing; in vitro developability assay.
Conflict of interest statement
The authors are current or past employees of Adimab, LLC and may hold shares in Adimab, LLC. K.B., A.S., and M.B.B. are authors on patents and patent applications describing heavy/light chain pairing mutations.
Figures



References
-
- The Antibody Society . Therapeutic monoclonal antibodies approved or in regulatory review [Internet]. [accessed 2024 Aug 1]. https://www.antibodysociety.org/antibody-therapeutics-product-data/.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
