Iterative hybrid model based optimization of rAAV production
- PMID: 40129076
- PMCID: PMC12348303
- DOI: 10.1002/btpr.70006
Iterative hybrid model based optimization of rAAV production
Abstract
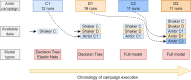
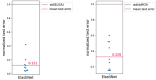
Changes in serotype or genetic payload of recombinant adeno associated virus (rAAVs) gene therapies require adapting the transfection conditions of the upstream HEK293 cultivations. This study adopts an iterative model-based experiment design approach, where increasing data availability is leveraged to evolve models of different complexity. Initial models based on data from shaker flask runs guided the design of the first round at Ambr250 scale. With Ambr250 data becoming available, hybrid models capturing process state evolutions and historical models incorporating these evolutions to predict rAAV titer, were developed. These models were then combined into a full model approach, which was utilized within a Bayesian Optimization framework for the design of a second round of Ambr250 scale runs. The iterative approach was tested across different projects applying transfer learning to enhance the predictive power and improve the subsequent optimization. The approach was benchmarked against a statistical Design of Experiment method. The results show that the model-based experiment design consistently (and across projects) produces higher rAAV titer values than the benchmark approach (Project C: 4.4% or 7.0% increases in titer values relative to the response surface modeling approach for ELISA and ddPCR, respectively; Project D: 32.4% or 10.9% increases in titer values relative to the standard DoE-screening pick for ELISA and ddPCR, respectively), effectively optimizing the transfection mixture composition. The combination of propagation and historical models, augmented by transfer learning and an ever-increasing amount of data, enhanced the process design workflow, contributing to improved rAAV production through efficient transfection strategies.
Keywords: Design of Experiments; Parallel Mini‐bioreactors; human embryonic kidney suspension cell; hybrid modeling; rAAV production.
© 2025 Baxalta Innovations GmbH, DataHow AG and The Author(s). Biotechnology Progress published by Wiley Periodicals LLC on behalf of American Institute of Chemical Engineers.
Conflict of interest statement
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Financial support was provided by Takeda and DataHow AG. All authors were employees of Takeda or Datahow AG at the time this study was performed.
Figures



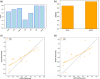





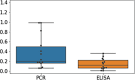
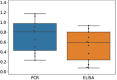
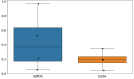
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

