Integration of Dynamical Network Biomarkers, Control Theory and Drosophila Model Identifies Vasa/DDX4 as the Potential Therapeutic Targets for Metabolic Syndrome
- PMID: 40136664
- PMCID: PMC11941168
- DOI: 10.3390/cells14060415
Integration of Dynamical Network Biomarkers, Control Theory and Drosophila Model Identifies Vasa/DDX4 as the Potential Therapeutic Targets for Metabolic Syndrome
Abstract
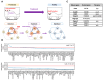
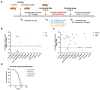
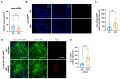
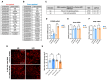
Metabolic syndrome (MetS) is a subclinical disease, resulting in increased risk of type 2 diabetes (T2D), cardiovascular diseases, cancer, and mortality. Dynamical network biomarkers (DNB) theory has been developed to provide early-warning signals of the disease state during a preclinical stage. To improve the efficiency of DNB analysis for the target genes discovery, the DNB intervention analysis based on the control theory has been proposed. However, its biological validation in a specific disease such as MetS remains unexplored. Herein, we identified eight candidate genes from adipose tissue of MetS model mice at the preclinical stage by the DNB intervention analysis. Using Drosophila, we conducted RNAi-mediated knockdown screening of these candidate genes and identified vasa (also known as DDX4), encoding a DEAD-box RNA helicase, as a fat metabolism-associated gene. Fat body-specific knockdown of vasa abrogated high-fat diet (HFD)-induced enhancement of starvation resistance through up-regulation of triglyceride lipase. We also confirmed that DDX4 expressing adipocytes are increased in HFD-fed mice and high BMI patients using the public datasets. These results prove the potential of the DNB intervention analysis to search the therapeutic targets for diseases at the preclinical stage.
Keywords: DNB intervention analysis; Drosophila melanogaster; dynamical network biomarkers theory; metabolic syndrome.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
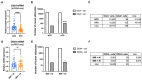
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

