Kaempferol Targets Global Epigenetic Modifiers to Impedes Growth and Migratory Ability of HeLa Cells
- PMID: 40159638
- PMCID: PMC11955415
- DOI: 10.1111/jcmm.70498
Kaempferol Targets Global Epigenetic Modifiers to Impedes Growth and Migratory Ability of HeLa Cells
Abstract
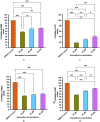
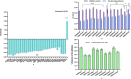
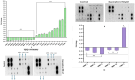
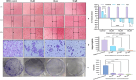
Dietary bioactive agents can curb tumour progression through chromatin alterations. Thus, this study attempts to evaluate the influence of kaempferol on epigenome modification in HeLa cells. Biochemical analysis for global DNA methylation-LINE 1, DNMTs (DNA methyltransferases), HAT (histone acetyl transferase), HDACs (histone deacetylases) and HMTs (histone methyltransferases) were examined with their transcript level expression through qPCR. Also, H3 and H4 histone modification marks were quantitated by an ELISA-based assay. Moreover, qPCR and protein profiler were performed to analyse the expression of migratory genes at both mRNA and protein levels, respectively, that was further substantiated through colony formation, invasion, and scratch wound assays. Finally, DNA methyl-sequencing was performed to analyse the promoter methylation of TSGs (tumour suppressor genes) and corroborated by analysing selected TSGs' expression. Kaempferol treatment did not alter the global DNA methylation-LINE 1 compared to untreated control, however, it reduced the expression and biochemical activities of DNMT and HDAC, which can be linked to their hypermethylation by kaempferol exposure. Concordant with the reduced expression of HMTs, HATs and other epi-enzymes, various histone H3 and H4 marks were also observed to be modulated. Kaempferol exposure led to promoter hypomethylation of various TSGs (such as WIF1, RUNX1, RARβ, SOX1), which subsequently led to enhanced expression at the mRNA level, which corresponds to their reactivation. Molecular studies were consistent with cell-based studies, which demonstrated a strong growth inhibitory and anti-migratory effect of kaempferol. This research helps to understand the probable mechanism used by kaempferol as a potential epigenetic modifier.
Keywords: epigenome; epi‐diet; hypermethylation; hypomethylation; kaempferol; methyl sequencing.
© 2025 The Author(s). Journal of Cellular and Molecular Medicine published by Foundation for Cellular and Molecular Medicine and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
References
-
- Wang L. H., Wu C. F., Rajasekaran N., and Shin Y. K., “Loss of Tumor Suppressor Gene Function in Human Cancer: An Overview,” Cellular Physiology and Biochemistry 51, no. 6 (2018): 2647–2693. - PubMed
-
- Walia Y. K. and Sharma V., “Role of HDACs and DNMTs in Cancer Therapy: A Review,” Asian Journal of Advanced Basic Science 1, no. 1 (2013): 62–78.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

