A genome-wide association study of imaging-defined atherosclerosis
- PMID: 40164586
- PMCID: PMC11958696
- DOI: 10.1038/s41467-025-57457-7
A genome-wide association study of imaging-defined atherosclerosis
Abstract
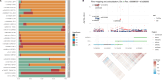
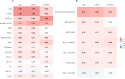
Imaging-defined atherosclerosis represents an intermediate phenotype of atherosclerotic cardiovascular disease (ASCVD). Genome-wide association studies (GWAS) on directly measured coronary plaques using coronary computed tomography angiography (CCTA) are scarce. In the so far largest population-based cohort with CCTA data, we performed a GWAS on coronary plaque burden as determined by the segment involvement score (SIS) in 24,811 European individuals. We identified 20 significant independent genetic markers for SIS, three of which were found in loci not implicated in ASCVD before. Further GWAS on coronary artery calcification showed similar results to that of SIS, whereas a GWAS on ultrasound-assessed carotid plaques identified both shared and non-shared loci with SIS. In two-sample Mendelian randomization studies using SIS-associated markers in UK Biobank and CARDIoGRAMplusC4D, one extra coronary segment with atherosclerosis corresponded to 1.8-fold increased odds of myocardial infarction. This GWAS data can aid future studies of causal pathways in ASCVD.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare the following competing interests: S.S. reports speakers’ honoraria and participation in the scientific board for the Exposure study (Janssen). The remaining authors declare no competing interests.
Figures



References
-
- Libby, P. Mechanisms of acute coronary syndromes and their implications for therapy. N. Engl. J. Med368, 2004–2013 (2013). - PubMed
-
- Erdmann, J., Kessler, T., Munoz Venegas, L. & Schunkert, H. A decade of genome-wide association studies for coronary artery disease: the challenges ahead. Cardiovasc Res114, 1241–1257 (2018). - PubMed
-
- Nelson, C. P. et al. Association analyses based on false discovery rate implicate new loci for coronary artery disease. Nat. Genet49, 1385–1391 (2017). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

