APOE4 impairs autophagy and Aβ clearance by microglial cells
- PMID: 40164781
- PMCID: PMC11958439
- DOI: 10.1007/s00011-025-02016-5
APOE4 impairs autophagy and Aβ clearance by microglial cells
Abstract
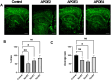
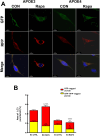
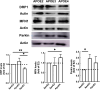
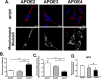
Alzheimer's disease (AD) is a predominant form of dementia in elderly. In sporadic AD and in families with higher risk of AD, correlation with apolipoprotein E4 (APOE) allele expression has been found. How APOE4 induces its pathological effects is still unclear. Several studies indicate that autophagy, a major degradation pathway trough the lysosome, may be compromised in AD. Here we studied, the effects of APOE isoforms expression in microglia cells. By using an in-situ model, the clearance of Aβ plaques from brain sections of transgenic 5xFAD mice by the APOE expressing microglia was examined. The results show that APOE4 microglia has Impairment In clearance of insoluble Aβ plaques as compared to APOE3 and APOE2 microglia. Furthermore, APOE4 affect the uptake of soluble Aβ. We found that microglia expressing APOE4 exhibit reduced autophagic flux as compared to those expressing APOE3. The autophagy inhibitor chloroquine also blocked Aβ plaque uptake in APOE3 expressing cells. Furthermore, we found that APOE4 expressing microglia have altered mitochondrial dynamics protein expression, mitochondrial morphology and mitochondrial activity compared to those expressing APOE2, and APOE3. Rapamycin treatment corrected Mitochondrial Membrane Potential in APOE4-expressing cells. Taken together, these findings suggest that APOE4 impairs the activation of autophagy, mitophagy, and Aβ clearance and that autophagy-inducing treatments, such as rapamycin, can enhance autophagy and mitochondrial functions in APOE4 expressing microglia. Our results reveal a direct link between APOE4 to autophagy activity in microglia, suggesting that the pathological effects of APOE4 could be counteracted by pharmacological treatments inducing autophagy, such as rapamycin.
Keywords: Alzheimer's disease (AD); Amyloid β; Apolipoprotein E4 (apoE4); Autophagy.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures





References
-
- Alzheimer A, Stelzmann RA, Schnitzlein HN, Murtagh FR. An english translation of Alzheimer’s 1907 paper, Uber eine eigenartige Erkankung der Hirnrinde. Clin Anat. 1995;8:429–31. - PubMed
-
- Masliah E, Crews L, Hansen L. Synaptic remodeling during aging and in Alzheimer’s disease. J Alzheimer’s Dis JAD. 2006;9:91–9. - PubMed
-
- Michaelson DM. APOE epsilon4: the most prevalent yet understudied risk factor for Alzheimer’s disease. Alzheimer’s Dement J Alzheimer’s Assoc. 2014;10:861–8. - PubMed
-
- Lee MJ, Lee JH, Rubinsztein DC. Tau degradation: the ubiquitin-proteasome system versus the autophagy-lysosome system. Prog Neurobiol. 2013;105:49–59. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

