TIMEPOINT, a phase 1 study combining MTL-CEBPA with pembrolizumab, supports the immunomodulatory effect of MTL-CEBPA in solid tumors
- PMID: 40168999
- PMCID: PMC12047497
- DOI: 10.1016/j.xcrm.2025.102041
TIMEPOINT, a phase 1 study combining MTL-CEBPA with pembrolizumab, supports the immunomodulatory effect of MTL-CEBPA in solid tumors
Abstract
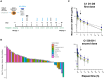
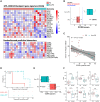
Many patients with cancer do not benefit from currently approved immune checkpoint inhibitors (ICIs), suggesting that additional immunomodulation of the immunosuppressive tumor microenvironment (TME) is required. MTL-CCAAT enhancer-binding protein alpha (CEBPA) specifically upregulates the expression of the master myeloid transcription factor, CEBPA, relieving myeloid-driven immunosuppression. Here, we report the safety, tolerability, pharmacokinetics, and efficacy of MTL-CEBPA in combination with pembrolizumab in patients with advanced solid tumors that typically show ICI resistance. Multimodal exploratory analyses of paired patient biopsies demonstrate biological changes associated with the combination treatment of MTL-CEBPA and pembrolizumab, including increased infiltration of T cell and antigen-presenting cells supporting conversion from an immune-desert toward a more immune-inflamed TME. Patients with disease stabilization demonstrate reductions in immunosuppressive myeloid cells post treatment. Collectively, these data support a role for MTL-CEBPA in reducing immunosuppression in the TME. This study was registered at ClinicalTrials.gov (NCT04105335).
Keywords: biomarker; cancer; gene activation; immunosuppression; immunotherapy; resistance; tumor microenvironment.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declarations of interests The following individuals are employees and shareholders of MiNA Therapeutics Ltd.: R. Habib, V.R., J.V., J.P.N., and N.A.H. M.H.S. has a research grant from MiNA Therapeutics Ltd. N.A.H. is the founder and director, and consults and advises at MiNA Therapeutics Ltd. and received travel, accommodation, and conference expenses support from MiNA Therapeutics Ltd. and institutional grant support from MiNA Therapeutics Ltd. A.E.-K. has received research support from Astex, has received personal fees from Merrimack, and has served as an advisor for Bristol Myers Squibb, AstraZeneca, Bayer, Genentech, and Novartis. T.M. has consultancies at Roche, AstraZeneca, Signant Health, GreyWolf, Guerbet, Geneos, Eisai, BeiGene, and MSD and received research funding from MSD, Bayer, and Boston Scientific. D.J.P. consults, advises, and is on the speakers’ bureau for Eisai and Roche and consults and advises for AstraZeneca, Avamune, BeiGene, Starpharma, BenevolentAI, Ipsen, and Mursla. He is on the speakers’ bureau and received grants from Bayer and Bristol Myers Squibb. He consults for Da Volterra, Exact, and Mina Therapeutics. He is on the speakers’ bureau for Falk Foundation and ViiV Healthcare. He received grants from GlaxoSmithKline, BMS, and MSD. Institutional Affiliation: secondary affiliation at the Department of Translational Medicine (DIMET), Universita' del Piemonte Orientale “A. Avogadro,” Novara, Italy. D. Sarker reports previous travel, accommodation, and conference expenses support from MiNA Therapeutics and receives consultancy or honoraria fees from Ipsen, Bayer, MSD, Roche, Servier, AstraZeneca, Boehringer, AbbVie, Sirtex, AAA, Incyte, and Eisai. B.K. has received consulting fees from Regeneron, travel funds from Genentech/Roche, and research funding (to institution) from MiNA Therapeutics Limited, Partner Therapeutics, Apexigen, Antengene, Innovative Cellular Therapeutics Co., AstraZeneca, Takeda, Regeneron, and Genentech/Roche. D.M. has received research funding from Amgen, Merck, Oncolytics, and Rafael; is on the scientific advisory board for Actuate and Qurient, is on an advisory/speaker bureau for Amgen, BMS, Eisai, and Exelixis; and has received funding paid to their institution from AbbVie, Acepodia, Actuate Therapeutics, ADC Therapeutics, Amgen, AVEO, Bayer, Blueprint Medicines, BMS, BioNTech, Dialectic Therapeutics, Epizyme, Fujifilm, ImmuneSensor, Immune-Onc Therapeutics, Leap Therapeutics, Lycera Corp., Merck, Millennium, MiNA Alpha, NGM Biopharmaceuticals, Novartis, Oncolytics, Orano Med, Puma, Qurient, Rafael, Repare Therapeutics, Triumvira Immunologics, Vigeo Therapeutics, and Werewolf Therapeutics. S.B. receives institutional research funding to conduct clinical trials (of which PI or CI) from NuCana PLC, UCB, BioNTech, Medannex, Nurix, Theolytics, and Regeneron; consults for Ellipses, Oxford Investment Consultants, UCB, Simbec-Orion, and Theolytics; is Director at RNA Guardian Ltd.; and holds an ownership interest. C.E.C. is an invited speaker at Amgen, AstraZeneca, Pierre Fabre, and Roche and is on the advisory board at Guardant Health AMEA, Merck, Pierre Fabre, and Roche. I.M. is on advisory boards for AstraZeneca, GSK, Clovis Oncology, pharma&, BioNTech, and Roche. T.R.J.E. declares honoraria (speaker’s fees; advisory boards) from MSD payable to his employing institution; support for investigator-led clinical trials (MSD, pembrolizumab) payable to his employing institution; and support to attend international conferences (MSD) – personal and is Editor-in-Chief of the British Journal of Cancer. M.S.N. receives research funding from Erytech and consults for Moderna and Merus. N.C. has received research grants from Roche Pharmaceuticals, consulting fees from Roche Pharmaceuticals and Redx Pharmaceuticals, payment or honoraria for lectures, presentations, speakers bureaus, manuscript writing, or educational events from Roche Pharmaceuticals, support for attending meetings and/or travel from Roche Pharmaceuticals, and research funding to research team from AstraZeneca, Orion, F. Hoffmann-La Roche Ltd., Taiho, GSK, Novartis, Starpharma, Bayer, Eisai, UCB, RedX Pharmaceuticals, Stemline Therapeutics, LOXO-oncology, Avacta, Boehringer Ingelheim, Merck, and Tarveda Therapeutics and participated in data safety monitoring board or advisory board at Roche Pharmaceuticals and Cancer Research UK.
Figures




References
-
- Robert C., Ribas A., Hamid O., Daud A., Wolchok J.D., Joshua A.M., Hwu W.J., Weber J.S., Gangadhar T.C., Joseph R.W., et al. Durable Complete Response After Discontinuation of Pembrolizumab in Patients With Metastatic Melanoma. J. Clin. Oncol. 2018;36:1668–1674. - PubMed
-
- Piha-Paul S.A., Oh D.Y., Ueno M., Malka D., Chung H.C., Nagrial A., Kelley R.K., Ros W., Italiano A., Nakagawa K., et al. Efficacy and safety of pembrolizumab for the treatment of advanced biliary cancer: Results from the KEYNOTE-158 and KEYNOTE-028 studies. Int. J. Cancer. 2020;147:2190–2198. - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Medical

