Asymmetric Enantio-complementary Synthesis of Thioethers via Ene-Reductase-Catalyzed C-C Bond Formation
- PMID: 40172483
- PMCID: PMC12147138
- DOI: 10.1021/jacs.5c00761
Asymmetric Enantio-complementary Synthesis of Thioethers via Ene-Reductase-Catalyzed C-C Bond Formation
Abstract
Enzymes are attractive catalysts due to their high chemo-, regio-, and enantioselectivity. In recent years, the application of enzymes in organic synthesis has expanded dramatically, especially for the synthesis of chiral alcohols and amines, two very important functional groups found in many active pharmaceutical ingredients (APIs). Indeed, many elegant routes employing such compounds have been described by industry. Yet, for the synthesis of chiral thiols and thioethers, likewise found in APIs albeit less ubiquitous, only very few biocatalytic syntheses have been reported, and stereocontrol has proved challenging. Here, we apply ene-reductases (EREDs), whose ability to initiate and control chemically challenging radical chemistries has recently emerged, to the synthesis of chiral thioethers from α-bromoacetophenones and pro-chiral vinyl sulfides, without requiring light. Depending on the choice of ERED either enantiomer of the product could be accessed. The highest conversion and selectivity were achieved with GluER T36A using fluorinated substrates, reaching up to 82% conversion and >99.5% ee. With α-bromoacetophenone and α-(methylthio)styrene, the reaction could be performed on a 100 mg scale, affording the product in a 46% isolated yield with a 93% ee. Finally, mechanistic studies were carried out using stopped-flow spectroscopy and protein mass spectrometry, providing insight into the preference of the enzyme for the intermolecular reaction. This work paves the way for new routes for the synthesis of thioether-containing compounds.
Figures

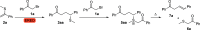




References
-
- Huffman M. A., Fryszkowska A., Alvizo O., Borra-Garske M., Campos K. R., Canada K. A., Devine P. N., Duan D., Forstater J. H., Grosser S. T., Halsey H. M., Hughes G. J., Jo J., Joyce L. A., Kolev J. N., Liang J., Maloney K. M., Mann B. F., Marshall N. M., McLaughlin M., Moore J. C., Murphy G. S., Nawrat C. C., Nazor J., Novick S., Patel N. R., Rodriguez-Granillo A., Robaire S. A., Sherer E. C., Truppo M. D., Whittaker A. M., Verma D., Xiao L., Xu Y., Yang H.. Design of an in Vitro Biocatalytic Cascade for the Manufacture of Islatravir. Science. 2019;366:1255–1259. doi: 10.1126/science.aay8484. - DOI - PubMed
-
- Prier C. K., Camacho Soto K., Forstater J. H., Kuhl N., Kuethe J. T., Cheung-Lee W. L., Di Maso M. J., Eberle C. M., Grosser S. T., Ho H. I., Hoyt E., Maguire A., Maloney K. M., Makarewicz A., McMullen J. P., Moore J. C., Murphy G. S., Narsimhan K., Pan W., Rivera N. R., Saha-Shah A., Thaisrivongs D. A., Verma D., Wyatt A., Zewge D.. Amination of a Green Solvent via Immobilized Biocatalysis for the Synthesis of Nemtabrutinib. ACS Catal. 2023;13:7707–7714. doi: 10.1021/acscatal.3c00941. - DOI
-
- Kumar R., Karmilowicz M. J., Burke D., Burns M. P., Clark L. A., Connor C. G., Cordi E., Do N. M., Doyle K. M., Hoagland S., Lewis C. A., Mangan D., Martinez C. A., Mcinturff E. L., Meldrum K., Pearson R., Steflik J., Rane A., Weaver J.. Biocatalytic Reductive Amination-Discovery to Commercial Manufacturing Applied to Abrocitinib JAK1 Inhibitor. Nat. Catal. 2021;4:775–782. doi: 10.21203/rs.3.rs-244144/v1. - DOI
-
- Schober M., MacDermaid C., Ollis A. A., Chang S., Khan D., Hosford J., Latham J., Ihnken L. A. F., Brown M. J. B., Fuerst D., Sanganee M. J., Roiban G. D.. Chiral Synthesis of LSD1 Inhibitor GSK2879552 Enabled by Directed Evolution of an Imine Reductase. Nat. Catal. 2019;2:909–915. doi: 10.1038/s41929-019-0341-4. - DOI
LinkOut - more resources
Full Text Sources

