HELLS controls mitochondrial dynamics and genome stability in liver cancer by collusion with MIEF1
- PMID: 40175344
- PMCID: PMC11965466
- DOI: 10.1038/s41419-025-07589-x
HELLS controls mitochondrial dynamics and genome stability in liver cancer by collusion with MIEF1
Abstract
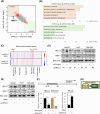
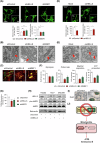
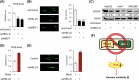
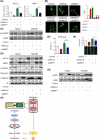
Dysregulated chromatin remodelers have emerged as critical disease targets. However, owing to the pleiotropic functions of chromatin remodelers, the underlying mechanisms of their effects on cancer have been difficult to elucidate. Here, we investigated the helicase lymphoid-specific (HELLS) oncogenic mechanism by identifying a new direct transcriptional target. Using loss or gain experiments, we identified Mitochondrial elongation factor 1 (MIEF1) as a critical target of the HELLS molecular network in liver cancer. Liver cancer patients with a poor prognosis exhibited upregulated expression of MIEF1, and MIEF1 knockdown led to the loss of tumor capabilities, indicating MIEF1 as an oncogene in liver cancer. Suppressing the HELLS-MIEF1 axis caused mitochondrial hyperfusion, energy deprivation, and further resulting senescence. HELLS knockdown globally increased histone 3 lysine 9 trimethylation (H3K9me3), especially in genomic hotspots with upregulation of SUV39H1 and further augmented DNA methylation. This stabilized genome and hyperfused mitochondria led to reduced levels of reactive oxygen species (ROS) and DNA damage. Finally, tumor cells became famished and calm. We further validated the functions of the HELLS-MIEF1 axis by MIEF1 overexpression and mitochondrial fusion drug. Our study has important implications for medical science by highlighting the crosstalk between epigenetics and metabolism through nuclear chromatin remodeler HELLS and mitochondrial protein MIEF1.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests. Ethics approval and consent to participate: This study was approved by the Institutional Animal Care and Use Committee of Konkuk University and was conducted in accordance with relevant guidelines and regulations.
Figures




References
-
- Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

