The inferred modulation of correlated vaginal microbiota and metabolome by cervical differentially expressed genes across distinct CIN grades
- PMID: 40175912
- PMCID: PMC11963661
- DOI: 10.1186/s12866-025-03922-8
The inferred modulation of correlated vaginal microbiota and metabolome by cervical differentially expressed genes across distinct CIN grades
Abstract
Background: In vitro studies have demonstrated the modulation of vaginal microbiota (VM) by cervical peptides which levels varied with the status of HPV infection and cervical intraepithelial neoplasia (CIN) grades. However, there is a deficiency in population-based studies investigating the modulation of VM compositions and metabolome by cervical differentially expressed genes (DEGs) across different grades of CIN.
Methods: This study included 43 HPV-positive women, classified into low-grade (CIN1, n = 23) and high-grade (CIN2 + , n = 20) groups. Vaginal swabs were collected for both microbiota and metabolome analysis. Cervical exfoliated cells were collected for RNA-Seq analysis.
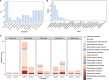
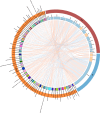
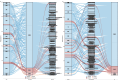
Results: We identified 258 differentially expressed genes (DEGs), among which 176 CIN1-enriched genes were linked to immune responses, cell chemotaxis, negative regulation of cell migration, and B cell differentiation, activation, and proliferation. Eighty-two genes upregulated in CIN2 + cohorts were associated with epidermis development and keratinization. Then, we identified 5,686 paired correlations between DEGs, VM, and metabolome, with 2,320 involving Lactobacillus. Further analysis revealed Lactobacillus as the primary determinant of metabolic profiles, followed by Gardnerella, Faecalibacterium, Aerococcus and Streptococcus, such as the notable positive correlation between Lactobacillus with D-lactic acid and DL-indole-3-lactic acid. Applying mediation analysis, we found that Lactobacillus mediated the association of 14 CIN1-enriched DEGs, such as COL4A2, CCBE1 and SPON1, with the production of 57 metabolites, including D-lactic acid, oleic acid and various amino acids. Additional analysis indicated significant mediation effects of 79 metabolites on the association of DEGs with the growth of Lactobacillus, Gardnerella, Fannyhessea and Aerococcus.
Conclusions: Our findings provide valuable population-based evidence for the inferred modulation of correlated VM and metabolome by cervical DEGs across different CIN stages.
Keywords: Cervical intraepithelial neoplasia; Host gene expression; Human papillomavirus; Vaginal metabolome; Vaginal microbiota.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: This study was approved by the Ethics Committee of Peking University Shenzhen Hospital (registration number: 2021–006). All participants were fully informed and then provided signed consents. We declare that our study adhered to the Declaration of Helsinki. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures

Similar articles
-
Inferred bi-directional interactions between vaginal microbiota, metabolome and persistent HPV infection accompanied by high-grade cervical intraepithelial neoplasia.BMC Microbiol. 2025 Jul 2;25(1):404. doi: 10.1186/s12866-025-04126-w. BMC Microbiol. 2025. PMID: 40604390 Free PMC article.
-
Microbial and metabolic profiles associated with HPV infection and cervical intraepithelial neoplasia: a multi-omics study.Microbiol Spectr. 2025 Jun 3;13(6):e0019225. doi: 10.1128/spectrum.00192-25. Epub 2025 Apr 30. Microbiol Spectr. 2025. PMID: 40304477 Free PMC article.
-
Microbiome and metabolomic changes associated with HPV clearance in women undergoing local excisional treatment for cervical intraepithelial neoplasia.mSystems. 2025 Jun 17;10(6):e0051125. doi: 10.1128/msystems.00511-25. Epub 2025 May 20. mSystems. 2025. PMID: 40391990 Free PMC article.
-
Associations of Atopobium, Garderella, Megasphaera, Prevotella, Sneathia, and Streptococcus with human papillomavirus infection, cervical intraepithelial neoplasia, and cancer: a systematic review and meta-analysis.BMC Infect Dis. 2025 May 16;25(1):708. doi: 10.1186/s12879-025-10851-4. BMC Infect Dis. 2025. PMID: 40380083 Free PMC article.
-
The Vaginal Microbiota, Human Papillomavirus, and Cervical Dysplasia-A Review.Medicina (Kaunas). 2025 May 5;61(5):847. doi: 10.3390/medicina61050847. Medicina (Kaunas). 2025. PMID: 40428805 Free PMC article. Review.
References
-
- de Sanjosé S, Brotons M, Pavón MA. The natural history of human papillomavirus infection. Best Pract Res Clin Obstet Gynaecol. 2018;47:2–13. 10.1016/j.bpobgyn.2017.08.015. - PubMed
-
- Delgado-Diaz DJ, Jesaveluk B, Hayward JA, Tyssen D, Alisoltani A, Potgieter M, Bell L, Ross E, Iranzadeh A, Allali I, Dabee S, Barnabas S, Gamieldien H, Blackburn JM, Mulder N, Smith SB, Edwards VL, Burgener AD, Bekker LG, Ravel J, Passmore JS, Masson L, Hearps AC, Tachedjian G. Lactic acid from vaginal microbiota enhances cervicovaginal epithelial barrier integrity by promoting tight junction protein expression. Microbiome. 2022;10(1):141. 10.1186/s40168-022-01337-5. - PMC - PubMed
-
- Berard AR, Brubaker DK, Birse K, Lamont A, Mackelprang RD, Noël-Romas L, Perner M, Hou X, Irungu E, Mugo N, Knodel S, Muwonge TR, Katabira E, Hughes SM, Levy C, Calienes FL, Lauffenburger DA, Baeten JM, Celum C, Hladik F, Lingappa J, Burgener AD. Vaginal epithelial dysfunction is mediated by the microbiome, metabolome, and mTOR signaling. Cell Rep. 2023;42(5):112474. 10.1016/j.celrep.2023.112474. - PMC - PubMed
-
- Nicolò S, Antonelli A, Tanturli M, Baccani I, Bonaiuto C, Castronovo G, Rossolini GM, Mattiuz G, Torcia MG. Bacterial species from vaginal microbiota differently affect the production of the E6 and E7 oncoproteins and of p53 and p-Rb oncosuppressors in HPV16-infected cells. Int J Mol Sci. 2023;24(8):7173. 10.3390/ijms24087173. - PMC - PubMed

