Identification of causative agents of infective endocarditis by metagenomic next-generation sequencing of resected valves
- PMID: 40182766
- PMCID: PMC11966046
- DOI: 10.3389/fcimb.2025.1532257
Identification of causative agents of infective endocarditis by metagenomic next-generation sequencing of resected valves
Abstract
Background: Infective endocarditis (IE) is a rare and life-threatening condition with considerable mortality rates. Diagnosis is often complicated by negative blood culture results, limiting the accurate identification of causative pathogens. This study aimed to evaluate the effectiveness of metagenomic next-generation sequencing (mNGS) of cardiac valve specimens compared to conventional clinical laboratory methods for identifying pathogens in IE.
Methods: Nineteen patients with suspected IE who were scheduled for surgical valve removal were prospectively enrolled. The metagenomic workflow included bacterial DNA enrichment from resected valves using the Molzym Ultra-Deep Microbiome Prep, sequencing of metagenomic libraries using the Illumina MiSeq platform, and Kraken 2 taxonomic assignments based on read data.
Results: Valve mNGS achieved a sensitivity of 82.4% and a specificity of 100% relative to the final adjudicated pathogen diagnosis. Blood culture, considered the reference standard, exhibited slightly higher sensitivity (88.2%) with comparable specificity (100%). In comparison, valve culture (sensitivity: 29.4%, specificity: 50.0%) and microscopy (sensitivity: 35.3%, specificity: 100%) showed lower diagnostic performance. Delays between blood culture negativization and valve resection impacted mNGS sensitivity, likely due to pathogen clearance. Notably, valves resected within 12 days from blood culture negativization achieved 100% diagnostic accuracy, emphasizing the importance of timing for optimal mNGS results.
Conclusion: This study underscores mNGS as a valuable diagnostic tool for detecting IE pathogens, complementing traditional diagnostic methods. The detection of antibiotic resistance genes and multi-locus sequence typing profiles in some samples further demonstrated its utility.
Keywords: cardiac surgery; clinical metagenomics; heart valve; microbiome; next-generation sequencing.
Copyright © 2025 Lazarevic, Gaïa, Pham, de Lorenzi-Tognon, Girard, Mauffrey, Charretier, Renzi, Huber and Schrenzel.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
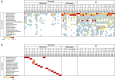

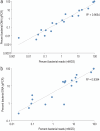
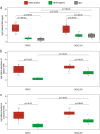

Similar articles
-
Detection of pathogens from resected heart valves of patients with infective endocarditis by next-generation sequencing.Int J Infect Dis. 2019 Jun;83:148-153. doi: 10.1016/j.ijid.2019.03.007. Epub 2019 Mar 27. Int J Infect Dis. 2019. PMID: 30926543
-
Identification of pathogens in culture-negative infective endocarditis cases by metagenomic analysis.Ann Clin Microbiol Antimicrob. 2018 Dec 20;17(1):43. doi: 10.1186/s12941-018-0294-5. Ann Clin Microbiol Antimicrob. 2018. PMID: 30567558 Free PMC article.
-
Application of Metagenomic Next-Generation Sequencing in the Etiological Diagnosis of Infective Endocarditis During the Perioperative Period of Cardiac Surgery: A Prospective Cohort Study.Front Cardiovasc Med. 2022 Mar 8;9:811492. doi: 10.3389/fcvm.2022.811492. eCollection 2022. Front Cardiovasc Med. 2022. PMID: 35369282 Free PMC article.
-
Infective endocarditis caused by Streptococcus sinensis diagnosed with next-generation sequencing: a case report and literature review.BMC Infect Dis. 2025 Mar 27;25(1):425. doi: 10.1186/s12879-025-10837-2. BMC Infect Dis. 2025. PMID: 40148753 Free PMC article. Review.
-
Unveiling the Future of Infective Endocarditis Diagnosis: The Transformative Role of Metagenomic Next-Generation Sequencing in Culture-Negative Cases.J Epidemiol Glob Health. 2025 Aug 22;15(1):108. doi: 10.1007/s44197-025-00455-1. J Epidemiol Glob Health. 2025. PMID: 40847264 Free PMC article. Review.
Cited by
-
Next-Generation Sequencing Techniques to Diagnose Culture-Negative Subacute Native Aortic Endocarditis.Emerg Infect Dis. 2025 Jul;31(7):1478-1481. doi: 10.3201/eid3107.241739. Emerg Infect Dis. 2025. PMID: 40562730 Free PMC article.
-
Early diagnosis of streptococcus cristatus in blood culture-negative infective endocarditis by capture-based metagenomic next-generation sequencing: a case report.Front Cardiovasc Med. 2025 Aug 7;12:1604687. doi: 10.3389/fcvm.2025.1604687. eCollection 2025. Front Cardiovasc Med. 2025. PMID: 40852206 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

