Clinical whole genome sequencing in pediatric epilepsy: Genetic and phenotypic spectrum of 733 individuals
- PMID: 40183601
- PMCID: PMC12371643
- DOI: 10.1111/epi.18403
Clinical whole genome sequencing in pediatric epilepsy: Genetic and phenotypic spectrum of 733 individuals
Abstract
Objective: A large proportion of pediatric epilepsies have an underlying genetic etiology. Limited studies have explored the efficacy of whole genome sequencing (WGS) in a clinical setting. Our academic-clinical center implemented clinical whole exome sequencing (WES) in 2014, then transitioned to WGS from 2015. We report the diagnostic yield, genetic and phenotypic findings, and prognostic factors following WGS/WES in pediatric epilepsy.
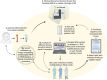
Methods: The cohort included 733 families with pediatric epilepsy who received clinical WGS/WES between 2014 and 2022. WGS/WES was performed at the Genomic Medicine Center Karolinska for Rare Diseases and analyzed at the Center for Inherited Metabolic Diseases at Karolinska University Hospital. Phenotypic information was extracted from referrals and medical records. Genetic and phenotypic data were analyzed using descriptive statistics, and univariable and multivariable analyses.
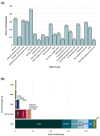
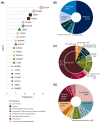
Results: The median age at seizure onset was 9 months. Developmental delay and/or intellectual disability (DD/ID) was observed in 61.3% of the cohort; 38.1% of individuals received an International League Against Epilepsy epilepsy syndrome diagnosis. WGS/WES was performed in 640 (87.3%) and 143 (19.5%) families, respectively, totaling 2029 individuals. A molecular diagnosis was identified in 278 of 733 individuals (37.9%), including 51 of 211 individuals analyzed more than once (24.2% of reanalyzed cases). Independent predictors for receiving a genetic diagnosis included female sex (adjusted odds ratio [aOR] = 1.8, 95% confidence interval [CI] = 1.3-2.4, p < .001), neonatal seizure onset (aOR = 2.5, 95% CI = 1.6-4, p < .001), mortality (aOR = 2.2, 95% CI = 1.3-4.0, p = .0048), and an ID/DD/developmental and epileptic encephalopathy (DEE) diagnosis (aOR = 1.8, 95% CI = 1.2-2.5, p = .0019). The strongest independent predictor of ID/DD/DEE was microcephaly (aOR = 7.8, 95% CI = 2-53, p = .0099). In the solved cohort, gene group did not predict cognitive outcome.
Significance: Clinical WGS is an effective diagnostic tool in pediatric epilepsy. We identified female sex as a novel prognostic factor for receiving a genetic diagnosis and highlight the value of reanalyzing previously unsolved cases to improve diagnostic yield.
Keywords: diagnostic yield; human phenotype ontology; infantile epilepsy; neurodevelopmental disorder; phenotypes; prognosis; real‐world data.
© 2025 The Author(s). Epilepsia published by Wiley Periodicals LLC on behalf of International League Against Epilepsy.
Conflict of interest statement
None of the authors has any conflict of interest to disclose. We confirm that we have read the Journal's position on issues involved in ethical publication and affirm that this report is consistent with those guidelines.
Figures

References
-
- Thomas RH, Berkovic SF. The hidden genetics of epilepsy‐a clinically important new paradigm. Nat Rev Neurol. 2014;10(5):283–292. - PubMed
-
- Lee HF, Chi CS, Tsai CR. Diagnostic yield and treatment impact of whole‐genome sequencing in paediatric neurological disorders. Dev Med Child Neurol. 2021;63(8):934–938. - PubMed
-
- Sheidley BR, Malinowski J, Bergner AL, Bier L, Gloss DS, Mu W, et al. Genetic testing for the epilepsies: a systematic review. Epilepsia. 2022;63(2):375–387. - PubMed
-
- Ferreira CR. The burden of rare diseases. Am J Med Genet A. 2019;179(6):885–892. - PubMed
-
- Hughes DA, Tunnage B, Yeo ST. Drugs for exceptionally rare diseases: do they deserve special status for funding? QJM. 2005;98(11):829–836. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

