A global collaboration for systematic analysis of broad-ranging antibodies against the SARS-CoV-2 spike protein
- PMID: 40184253
- PMCID: PMC12014896
- DOI: 10.1016/j.celrep.2025.115499
A global collaboration for systematic analysis of broad-ranging antibodies against the SARS-CoV-2 spike protein
Abstract
The Coronavirus Immunotherapeutic Consortium (CoVIC) conducted side-by-side comparisons of over 400 anti-SARS-CoV-2 spike therapeutic antibody candidates contributed by large and small companies as well as academic groups on multiple continents. Nine reference labs analyzed antibody features, including in vivo protection in a mouse model of infection, spike protein affinity, high-resolution epitope binning, ACE-2 binding blockage, structures, and neutralization of pseudovirus and authentic virus infection, to build a publicly accessible dataset in the database CoVIC-DB. High-throughput, high-resolution binning of CoVIC antibodies defines a broad and predictive landscape of antibody epitopes on the SARS-CoV-2 spike protein and identifies features associated with durable potency against multiple SARS-CoV-2 variants of concern and high in vivo efficacy. Results of the CoVIC studies provide a guide for selecting effective and durable antibody therapeutics and for immunogen design as well as providing a framework for rapid response to future viral disease outbreaks.
Keywords: COVID-19; CP: Immunology; SARS-CoV-2 spike protein; coronavirus; high-resolution epitope binning; neutralization; therapeutic antibodies.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests R.S.B. is a member the VaxArt, Takeda, and Invivyd advisory boards and has collaborative projects with Gilead, J&J, and HilleVax that are unrelated to this work. R.S.B. is a co-inventor of methods and uses of mouse-adapted and derivative SARS-CoV-2 viruses (US patent US11225508B1). D.B. and T.G. are employees of Carterra. Y.K. has received unrelated funding support from Daiichi Sankyo Pharmaceutical; Toyama Chemical; Tauns Laboratories, Inc.; Shionogi & Co. Ltd.; Otsuka Pharmaceutical; KM Biologics; Kyoritsu Seiyaku; Shinya Corporation; and Fuji Rebio.
Figures






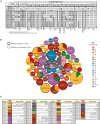

References
-
- WHO Coronavirus (COVID-19) dashboard https://covid19.who.int/.
-
- Bamlanivimab EUA letter of authorization https://www.fda.gov/media/143602.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

