This is a preprint.
A method for the detection and enrichment of endogenous cereblon substrates
- PMID: 40196695
- PMCID: PMC11974815
- DOI: 10.1101/2025.03.24.645063
A method for the detection and enrichment of endogenous cereblon substrates
Update in
-
A method for the detection and enrichment of endogenous cereblon substrates.Cell Chem Biol. 2025 Aug 21;32(8):1028-1041.e13. doi: 10.1016/j.chembiol.2025.07.002. Epub 2025 Aug 8. Cell Chem Biol. 2025. PMID: 40782806
Abstract
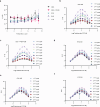
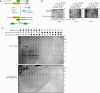
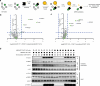
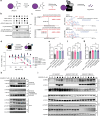
C-Terminal cyclic imides are posttranslational modifications on proteins that are recognized and removed by the E3 ligase substrate adapter cereblon (CRBN). Despite the observation of these modifications across the proteome by mass spectrometry-based proteomics, an orthogonal and generalizable method to visualize the C-terminal cyclic imide would enhance detection, sensitivity, and throughput of endogenous CRBN substrate characterization. Here we develop an antibody-like reagent, termed "cerebody," for visualizing and enriching C-terminal cyclic imide-modified proteins. We describe the engineering of CRBN derivatives to produce cerebody and use it to identify CRBN substrates by Western blot and enrichment from whole cell and tissue lysates. CRBN substrates identified by cerebody enrichment are mapped, validated, and further characterized for dependence on the C-terminal cyclic imide modification. These methods will accelerate the characterization of endogenous CRBN substrates and their regulation.
Conflict of interest statement
Declaration of Interest R.M. and N.C.P. are inventors on patent applications related to the CoraFluor TR-FRET technology used in this work. The Woo Lab receives or has received sponsored research support from Amgen, Ono Pharmaceuticals, and Merck. The Ciulli laboratory receives or has received sponsored research support from Almirall, Amgen, Amphista Therapeutics, Boehringer Ingelheim, GlaxoSmithKline, Eisai, Merck KGaA, Nurix Therapeutics, Ono Pharmaceuticals, and Tocris-Biotechne. A.C. is a co-founder and shareholder of Amphista Therapeutics, a company that is developing targeted protein degradation therapeutic platforms.
Figures







Similar articles
-
A method for the detection and enrichment of endogenous cereblon substrates.Cell Chem Biol. 2025 Aug 21;32(8):1028-1041.e13. doi: 10.1016/j.chembiol.2025.07.002. Epub 2025 Aug 8. Cell Chem Biol. 2025. PMID: 40782806
-
Genesis and regulation of C-terminal cyclic imides from protein damage.Proc Natl Acad Sci U S A. 2025 Jan 7;122(1):e2415976121. doi: 10.1073/pnas.2415976121. Epub 2024 Dec 30. Proc Natl Acad Sci U S A. 2025. PMID: 39793072 Free PMC article.
-
The contribution of cyclic imide stereoisomers on cereblon-dependent activity.Chem Sci. 2025 May 28;16(25):11519-11529. doi: 10.1039/d5sc01371b. eCollection 2025 Jun 25. Chem Sci. 2025. PMID: 40443985 Free PMC article.
-
Interventions to reduce harm from continued tobacco use.Cochrane Database Syst Rev. 2016 Oct 13;10(10):CD005231. doi: 10.1002/14651858.CD005231.pub3. Cochrane Database Syst Rev. 2016. PMID: 27734465 Free PMC article.
-
Automated devices for identifying peripheral arterial disease in people with leg ulceration: an evidence synthesis and cost-effectiveness analysis.Health Technol Assess. 2024 Aug;28(37):1-158. doi: 10.3310/TWCG3912. Health Technol Assess. 2024. PMID: 39186036 Free PMC article.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
