The stage- and subgroup-specific impact of non-HLA polymorphisms on preclinical type 1 diabetes progression
- PMID: 40196768
- PMCID: PMC11947702
- DOI: 10.1016/j.heliyon.2025.e42156
The stage- and subgroup-specific impact of non-HLA polymorphisms on preclinical type 1 diabetes progression
Abstract
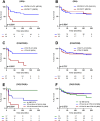
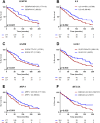
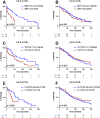
Besides variation within the HLA gene complex determining a major part of genetic susceptibility to Type 1 diabetes, genome-wide association studies have identified over 60 non-HLA loci also contributing to disease risk. While individual single nucleotide polymorphisms (SNPs) have limited predictive power, genetic risk scores (GRS) can identify at-risk individuals. However, current models do not fully capture the heterogeneous progression of asymptomatic islet autoimmunity, especially in autoantibody-positive subjects. In this study, we investigated the additional stage-specific impact of 17 non-HLA loci on previously established prediction models in 448 persistently autoantibody-positive first-degree relatives. Cox regression and Kaplan Meier survival analysis were used to assess their influence on progression from single to multiple autoantibody-positivity, and from there to clinical onset. FUT2 and CTSH significantly accelerated progression of single to multiple autoAb-positivity, but only in presence of insulin autoantibodies and HLA-DQ2/DQ8, respectively. At the stage of multiple autoantibody-positivity, progression to clinical onset was impacted by various non-HLA SNPs either as independent predictors (GLIS3, CENPW, IL2, GSDM, MEG3A, and NRP-1) or through interaction with HLA class I alleles (CLEC16A, NRP-1, TCF7L2), maternal diabetes status (CTSH), or a high-risk autoantibody-profile (CD226). Our data indicate that, unlike for GRS, the weight of distinct non-HLA polymorphisms varies significantly among individuals at risk, depending on disease stage and other stage-specific risk factors. They refine our previous stage-specific prediction models including age, autoantibody-profile, HLA genotype, and other non-HLA SNPs, and emphasize the importance of stratifying accordingly to personalize time-to-event prediction in risk groups, or for preparing or interpreting prevention trials.
Keywords: Autoimmunity; Non-HLA SNPs; Prediabetes; Prediction; Type 1 diabetes.
© 2025 The Author(s).
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures
Similar articles
-
CTLA4, SH2B3, and CLEC16A diversely affect the progression of early islet autoimmunity in relatives of Type 1 diabetes patients.Clin Exp Immunol. 2023 Mar 24;211(3):224-232. doi: 10.1093/cei/uxad002. Clin Exp Immunol. 2023. PMID: 36622793 Free PMC article.
-
Twenty-Year Progression Rate to Clinical Onset According to Autoantibody Profile, Age, and HLA-DQ Genotype in a Registry-Based Group of Children and Adults With a First-Degree Relative With Type 1 Diabetes.Diabetes Care. 2017 Aug;40(8):1065-1072. doi: 10.2337/dc16-2228. Diabetes Care. 2017. PMID: 28701370
-
Accelerated Progression to Type 1 Diabetes in the Presence of HLA-A*24 and -B*18 Is Restricted to Multiple Islet Autoantibody-Positive Individuals With Distinct HLA-DQ and Autoantibody Risk Profiles.Diabetes Care. 2018 May;41(5):1076-1083. doi: 10.2337/dc17-2462. Epub 2018 Mar 15. Diabetes Care. 2018. PMID: 29545461
-
Genetic prediction of autoimmunity: initial oligogenic prediction of anti-islet autoimmunity amongst DR3/DR4-DQ8 relatives of patients with type 1A diabetes.J Autoimmun. 2005;25 Suppl:40-5. doi: 10.1016/j.jaut.2005.09.002. Epub 2005 Oct 19. J Autoimmun. 2005. PMID: 16242305 Review.
-
Risk of beta-cell autoimmunity presence for progression to type 1 diabetes: A systematic review and meta-analysis.J Autoimmun. 2018 Jan;86:9-18. doi: 10.1016/j.jaut.2017.09.012. Epub 2017 Oct 5. J Autoimmun. 2018. PMID: 28988642
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

