Edge-enhanced interaction graph network for protein-ligand binding affinity prediction
- PMID: 40198678
- PMCID: PMC11977954
- DOI: 10.1371/journal.pone.0320465
Edge-enhanced interaction graph network for protein-ligand binding affinity prediction
Abstract
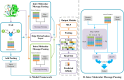
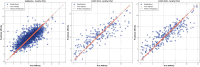
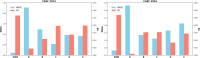
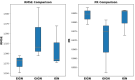
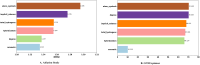
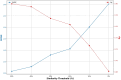
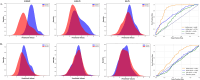
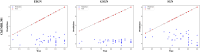
Protein-ligand interactions are crucial in drug discovery. Accurately predicting protein-ligand binding affinity is essential for screening potential drugs. Graph neural networks have proven highly effective in modeling spatial relationships and three-dimensional structures within intermolecular. In this paper, we introduce a graph neural network-based model named EIGN to predict protein-ligand binding affinity. The model consists of three main components: the normalized adaptive encoder, the molecular information propagation module, and the output module. Experimental results indicate that EIGN achieves root mean squared error of 1.126 and Pearson correlation coefficient of 0.861 on CASF-2016. Additionally, our model outperforms state-of-the-art methods on CASF-2013, CASF-2016, and the CSAR-NRC set, showing exceptional accuracy and robust generalization ability. To further validate the effectiveness of EIGN, we conducted several experiments, including ablation studies, feature importance analysis, data similarity analysis, and others, to evaluate its performance and applicability.
Copyright: © 2025 Yang et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Similar articles
-
PLAGCA: Predicting protein-ligand binding affinity with the graph cross-attention mechanism.J Biomed Inform. 2025 May;165:104816. doi: 10.1016/j.jbi.2025.104816. Epub 2025 Mar 24. J Biomed Inform. 2025. PMID: 40139623
-
Predicting Protein-Ligand Binding Affinity Using Fusion Model of Spatial-Temporal Graph Neural Network and 3D Structure-Based Complex Graph.Interdiscip Sci. 2025 Jun;17(2):257-276. doi: 10.1007/s12539-024-00644-9. Epub 2024 Nov 14. Interdiscip Sci. 2025. PMID: 39541085
-
Enhancing protein-ligand binding affinity prediction through sequential fusion of graph and convolutional neural networks.J Comput Chem. 2024 Dec 15;45(32):2929-2940. doi: 10.1002/jcc.27499. Epub 2024 Sep 2. J Comput Chem. 2024. PMID: 39223071
-
Predicting Protein-Ligand Docking Structure with Graph Neural Network.J Chem Inf Model. 2022 Jun 27;62(12):2923-2932. doi: 10.1021/acs.jcim.2c00127. Epub 2022 Jun 14. J Chem Inf Model. 2022. PMID: 35699430 Free PMC article. Review.
-
Advances in Protein-Ligand Binding Affinity Prediction via Deep Learning: A Comprehensive Study of Datasets, Data Preprocessing Techniques, and Model Architectures.Curr Drug Targets. 2024;25(15):1041-1065. doi: 10.2174/0113894501330963240905083020. Curr Drug Targets. 2024. PMID: 39318214 Free PMC article. Review.
References
-
- Metz J, Hajduk P. Rational approaches to targeted polypharmacology: Creating and navigating protein–ligand interaction networks. Curr Opin Chem Biol. 2010;14(4):498–504. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

