PACT prevents aberrant activation of PKR by endogenous dsRNA without sequestration
- PMID: 40199855
- PMCID: PMC11978871
- DOI: 10.1038/s41467-025-58433-x
PACT prevents aberrant activation of PKR by endogenous dsRNA without sequestration
Abstract
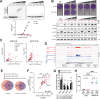
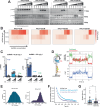
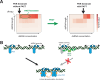
The innate immune sensor PKR for double-stranded RNA (dsRNA) is critical for antiviral defense, but its aberrant activation by cellular dsRNA is linked to various diseases. The dsRNA-binding protein PACT plays a critical yet controversial role in this pathway. We show that PACT directly suppresses PKR activation by endogenous dsRNA ligands, such as inverted-repeat Alu RNAs, which robustly activate PKR in the absence of PACT. Instead of competing for dsRNA binding, PACT prevents PKR from scanning along dsRNA-a necessary step for PKR molecules to encounter and phosphorylate each other for activation. While PKR favors longer dsRNA for increased co-occupancy and scanning-mediated activation, longer dsRNA is also more susceptible to PACT-mediated regulation due to increased PACT-PKR co-occupancy. Unlike viral inhibitors that constitutively suppress PKR, this RNA-dependent mechanism allows PACT to fine-tune PKR activation based on dsRNA length and quantity, ensuring self-tolerance without sequestering most cellular dsRNA.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




References
-
- Kim, Y. et al. PKR senses nuclear and mitochondrial signals by interacting with endogenous double-stranded RNAs. Mol. Cell71, 1051–1063.e1056 (2018). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

