A beginner's approach to deep learning applied to VS and MD techniques
- PMID: 40200329
- PMCID: PMC11980327
- DOI: 10.1186/s13321-025-00985-7
A beginner's approach to deep learning applied to VS and MD techniques
Abstract
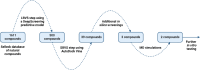
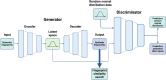
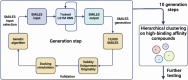
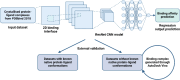
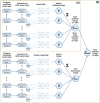
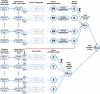
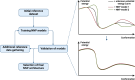
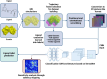
It has become impossible to imagine the fields of biochemistry and medicinal chemistry without computational chemistry and molecular modelling techniques. In many steps of the drug development process in silico methods have become indispensable. Virtual screening (VS) can tremendously expedite the early discovery phase, whilst the use of molecular dynamics (MD) simulations forms a powerful additional tool to in vitro methods throughout the entire drug discovery process. In the field of biochemistry, MD has also become a compelling method for studying biophysical systems (e.g., protein folding) complementary to experimental techniques. However, both VS and MD come with their own limitations and methodological difficulties, from hardware limitations to restrictions in algorithmic capabilities. One solution to overcoming these difficulties lies in the field of machine learning (ML), and more specifically deep learning (DL). There are many ways in which DL can be applied to these molecular modelling techniques to achieve more accurate results in a more efficient manner or expedite the data analysis of the acquired results. Despite steadily increasing interest in DL amidst computational chemists, knowledge is still limited and scattered over different resources. This review is aimed at computational chemists with knowledge of molecular modelling, who wish to possibly integrate DL approaches in their research and already have a basic understanding of the fundamentals of DL. This review focusses on a survey of recent applications of DL in molecular modelling techniques. The different sections are logically subdivided, based on where DL is integrated in the research: (1) for the improvement of VS workflows, (2) for the improvement of certain workflows in MD simulations, (3) for aiding in the calculations of interatomic forces, or (4) for data analysis of MD trajectories. It will become clear that DL has the capacity to completely transform the way molecular modelling is carried out.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures


Similar articles
-
Trends in Deep Learning for Property-driven Drug Design.Curr Med Chem. 2021;28(38):7862-7886. doi: 10.2174/0929867328666210729115728. Curr Med Chem. 2021. PMID: 34325627
-
Deep learning tools for advancing drug discovery and development.3 Biotech. 2022 May;12(5):110. doi: 10.1007/s13205-022-03165-8. Epub 2022 Apr 9. 3 Biotech. 2022. PMID: 35433167 Free PMC article. Review.
-
Molecular dynamics and quantum mechanics of RNA: conformational and chemical change we can believe in.Acc Chem Res. 2010 Jan 19;43(1):40-7. doi: 10.1021/ar900093g. Acc Chem Res. 2010. PMID: 19754142 Free PMC article.
-
Operando Modeling of Zeolite-Catalyzed Reactions Using First-Principles Molecular Dynamics Simulations.ACS Catal. 2023 Aug 15;13(17):11455-11493. doi: 10.1021/acscatal.3c01945. eCollection 2023 Sep 1. ACS Catal. 2023. PMID: 37671178 Free PMC article. Review.
-
Molecular Dynamics Simulations Are Redefining Our View of Peptides Interacting with Biological Membranes.Acc Chem Res. 2018 May 15;51(5):1106-1116. doi: 10.1021/acs.accounts.7b00613. Epub 2018 Apr 18. Acc Chem Res. 2018. PMID: 29667836
Cited by
-
Editorial: Exploring molecular recognition: integrating experimental and computational approaches.Front Mol Biosci. 2025 Jun 10;12:1637793. doi: 10.3389/fmolb.2025.1637793. eCollection 2025. Front Mol Biosci. 2025. PMID: 40556692 Free PMC article. No abstract available.
References
-
- De Vivo M, Masetti M, Bottegoni G, Cavalli A (2016) Role of molecular dynamics and related methods in drug discovery. J Med Chem 59:4035–4061. 10.1021/ACS.JMEDCHEM.5B01684 - PubMed
-
- Amini A, Amini A, Lolla S. (2024) MIT 6.S191 | Introduction to deep learning. http://introtodeeplearning.com/.
-
- Abadi M, Agarwal A, Barham P, Brevdo E, Chen Z, Citro C et al (2016) TensorFlow: large-scale machine learning on heterogeneous distributed systems. ArXiv. 10.48550/arXiv.1603.04467
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

