Vertical DNA Nanostructure Arrays: Facilitating Functionalization on Macro-Scale Surfaces
- PMID: 40200829
- PMCID: PMC12120987
- DOI: 10.1021/acsnano.5c03100
Vertical DNA Nanostructure Arrays: Facilitating Functionalization on Macro-Scale Surfaces
Abstract
The capability for varied functionalization and precise control at the nanoscale are significant advantages of DNA nanostructures. In the assembly of DNA nanostructure, the surface-assisted growth method utilizing double-crossover (DX) tile structures facilitates nucleation at relatively low concentrations on the surface based on electrostatic interactions, thereby enabling crystal growth over large areas. However, in surface-assisted growth, the geometrical hindrance of vertical structures on the DX tile structure surface makes it challenging to conjugate DNA nanostructures into fabricated surfaces. Here, the surface-assisted growth method was employed to extend the DX tile growth for forming vertical structure arrays on the substrate, providing attachment sites for functionalization on uniformly covered substrates at the macroscopic scale. Additionally, the spacing of the vertical structure arrays was demonstrated to be controllable through the strategic design of the repeating unit tiles that construct the DX crystals.
Keywords: AFM; DNA nanotechnology; aptamer; double-crossover tile; macro-scale; supporting substrate.
Figures



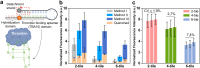
References
-
- Xu X. Z., Zhang Z. H., Qiu L., Zhuang J. N., Zhang L., Wang H., Liao C. N., Song H. D., Qiao R. X., Gao P., Hu Z. H., Liao L., Liao Z. M., Yu D. P., Wang E. G., Ding F., Peng H. L., Liu K. H.. Ultrafast Growth of Single-Crystal Graphene Assisted by a Continuous Oxygen Supply. Nat. Nanotechnol. 2016;11:930–935. doi: 10.1038/nnano.2016.132. - DOI - PubMed
-
- Shepelenko M., Hirsch A., Varsano N., Beghi F., Addadi L., Kronik L., Leiserowitz L.. Polymorphism, Structure, and Nucleation of Cholesterol·H2o at Aqueous Interfaces and in Pathological Media: Revisited from a Computational Perspective. J. Am. Chem. Soc. 2022;144:5304–5314. doi: 10.1021/jacs.1c10563. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

