Establishment of patient-derived xenograft models in Chinese patients with multiple myeloma: Insights into therapeutic responsiveness and molecular subtyping
- PMID: 40203791
- PMCID: PMC12011171
- DOI: 10.1016/j.tranon.2025.102385
Establishment of patient-derived xenograft models in Chinese patients with multiple myeloma: Insights into therapeutic responsiveness and molecular subtyping
Abstract
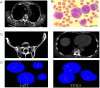
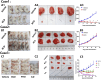
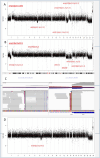
Multiple myeloma (MM), a malignant hematologic tumor characterized by the proliferation of monoclonal plasma cells, remains incurable with high relapse rates despite advances in treatment. Patient-derived xenograft (PDX) models have emerged as a promising tool for understanding MM's complex pathophysiology and testing therapeutic responses. In this study, we successfully developed PDX models from three patients with MM by subcutaneously engrafting their tumor cells into immunodeficient NCG mice. These models accurately mirrored the clinical drug responses of their corresponding patient cases, exhibiting similar drug sensitivities and resistance patterns. Omics profiling facilitated the alignment of PDX models with specific molecular subgroups identified in current MM research, enhancing the models' clinical relevance. The concordance between PDX models and clinical data confirms the utility of these models in simulating patient-specific responses and advancing personalized treatment strategies. This study validates the effectiveness of PDX models established by subcutaneous engraftment of tumor cells in replicating human disease and treatment responses, thus providing a robust platform for future personalized treatments and development of targeted interventions in Chinese MM patients.
Keywords: Drug responsiveness; Molecular subgroups therapeutic innovations; Multiple myeloma; Omics profiling; Patient-derived xenograft; Subcutaneous engraftment.
Copyright © 2025. Published by Elsevier Inc.
Conflict of interest statement
Declaration of competing interest Danyi Wen is an employee of LIDE Biotech and provided us with technical guidance for PDX model but have no conflict of interest.
Figures

Similar articles
-
Patient-Derived Xenograft Models for Intrahepatic Cholangiocarcinoma and Their Application in Guiding Personalized Medicine.Front Oncol. 2021 Jul 13;11:704042. doi: 10.3389/fonc.2021.704042. eCollection 2021. Front Oncol. 2021. PMID: 34327143 Free PMC article.
-
Characterization of drug responses of mini patient-derived xenografts in mice for predicting cancer patient clinical therapeutic response.Cancer Commun (Lond). 2018 Sep 26;38(1):60. doi: 10.1186/s40880-018-0329-5. Cancer Commun (Lond). 2018. PMID: 30257718 Free PMC article.
-
Patient-Derived Tumor Xenograft Models: Toward the Establishment of Precision Cancer Medicine.J Pers Med. 2020 Jul 18;10(3):64. doi: 10.3390/jpm10030064. J Pers Med. 2020. PMID: 32708458 Free PMC article. Review.
-
Establishment of a high-fidelity patient-derived xenograft model for cervical cancer enables the evaluation of patient's response to conventional and novel therapies.J Transl Med. 2023 Sep 9;21(1):611. doi: 10.1186/s12967-023-04444-5. J Transl Med. 2023. PMID: 37689699 Free PMC article.
-
Patient-derived xenograft (PDX) models: characteristics and points to consider for the process of establishment.J Toxicol Pathol. 2020 Jul;33(3):153-160. doi: 10.1293/tox.2020-0007. Epub 2020 Mar 20. J Toxicol Pathol. 2020. PMID: 32764840 Free PMC article. Review.
References
-
- van de Donk N., Pawlyn C., Yong K.L. Multiple myeloma. Lancet. 2021;397(10272):410–427. - PubMed
-
- Cowan A.J., Green D.J., Kwok M., Lee S., Coffey D.G., Holmberg L.A., Tuazon S., Gopal A.K., Libby E.N. Diagnosis and management of multiple myeloma: a review. JAMa. 2022;327(5):464–477. - PubMed
-
- Siegel R.L., Miller K.D., Jemal A. Cancer statistics, 2020. CA Cancer J. Clin. 2020;70(1):7–30. - PubMed
LinkOut - more resources
Full Text Sources

