Integrative biophysical characterization of molecular interactions: A case study with the sfGFP-nanobody complex
- PMID: 40204056
- PMCID: PMC12131814
- DOI: 10.1016/j.ab.2025.115859
Integrative biophysical characterization of molecular interactions: A case study with the sfGFP-nanobody complex
Abstract
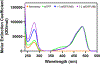
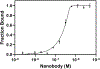
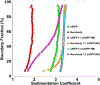
This study compares several analytical biophysical methods for investigating protein-protein interactions (PPIs) in solution, using the interaction between superfolder green fluorescent protein (sfGFP) and its anti-sfGFP nanobody enhancer as a model system. Techniques evaluated include microscale thermophoresis, fluorescence correlation spectroscopy, analytical ultracentrifugation with multi-wavelength and fluorescence detection, isothermal titration calorimetry, and analytical size exclusion chromatography coupled to multi-angle static light scattering and dynamic light scattering. Each method was assessed for information content, dynamic range, precision, and complementarity. The results consistently indicate a single-digit nanomolar dissociation constant and 1:1 stoichiometry for the interaction. While each technique offers unique insights into binding affinity, thermodynamics, and stoichiometry of the interaction, the multi-method approach provides a more complete and reliable characterization of PPIs. The study demonstrates how combining multiple complementary techniques enhances the robustness of PPI analysis in solution-phase conditions.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of competing interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures
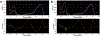

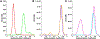
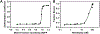
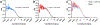
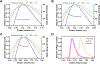
Similar articles
-
Screening approaches for the identification of Nrf2-Keap1 protein-protein interaction inhibitors targeting hot spot residues.SLAS Discov. 2024 Mar;29(2):100125. doi: 10.1016/j.slasd.2023.11.001. Epub 2023 Nov 5. SLAS Discov. 2024. PMID: 37935317
-
Complexity associated with caprylate binding to bovine serum albumin: Dimerization, allostery, and variance between the change in free energy and enthalpy of binding.Protein Sci. 2024 Dec;33(12):e5224. doi: 10.1002/pro.5224. Protein Sci. 2024. PMID: 39548833
-
[Preparation and chromatographic performance evaluation of hydrophilic interaction chromatography stationary phase based on amino acids].Se Pu. 2025 Jul;43(7):734-743. doi: 10.3724/SP.J.1123.2025.04015. Se Pu. 2025. PMID: 40610768 Free PMC article. Chinese.
-
Quality improvement strategies for diabetes care: Effects on outcomes for adults living with diabetes.Cochrane Database Syst Rev. 2023 May 31;5(5):CD014513. doi: 10.1002/14651858.CD014513. Cochrane Database Syst Rev. 2023. PMID: 37254718 Free PMC article.
-
Management of urinary stones by experts in stone disease (ESD 2025).Arch Ital Urol Androl. 2025 Jun 30;97(2):14085. doi: 10.4081/aiua.2025.14085. Epub 2025 Jun 30. Arch Ital Urol Androl. 2025. PMID: 40583613 Review.
References
-
- Kirchhofer A, Helma J, Schmidthals K, Frauer C, Cui S, Karcher A, Pellis M, Muyldermans S, Casas-Delucchi CS, Cardoso MC, Leonhardt H, Hopfner KP, Rothbauer U, Modulation of protein properties in living cells using nanobodies, Nat. Struct. Mol. Biol. 17 (1) (2010. Jan) 133–138, 10.1038/nsmb.1727. Epub 2009 Dec 13. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

