Salt stress alters the selectivity of mature pecan for the rhizosphere community and its associated functional traits
- PMID: 40206877
- PMCID: PMC11979281
- DOI: 10.3389/fpls.2025.1473473
Salt stress alters the selectivity of mature pecan for the rhizosphere community and its associated functional traits
Abstract
Introduction: Salt stress is a major global environmental factor limiting plant growth. Rhizosphere bacteria, recruited from bulk soil, play a pivotal role in enhancing salt stress resistance in herbaceous and crop species. However, whether the rhizosphere bacterial community of a mature tree can respond to salt stress, particularly in saline-alkalitolerant trees, remains unexplored. Pecan (Carya illinoinensis), an important commercially cultivated nut tree, is considered saline-alkali tolerant.
Methods: Pecan trees (12 years) were subjected to different NaCl concentrations for 12 weeks. Collected samples included bulk soil, rhizosphere soil, roots, leaves, and fruit. Amplicon sequencing data and shotgun metagenomic sequencing data obtained from the samples were investigated: 1) microbial communities in various ecological niches of mature pecan trees; 2) the characteristic of the rhizosphere bacteria community and the associated functional traits when pecan suffered from salt stress.
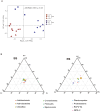
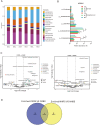
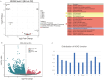
Results and discussion: We characterized the mature pecan-associated microbiome (i.e., fruit, leaf, root, and rhizosphere soil) for the first time. These findings suggest that niche-based processes, such as habitat selection, drive bacterial and fungal community assembly in pecan tissues. Salt stress reduced bacterial diversity, altered community composition, and shifted pecan's selective pressure on Proteobacteria and Actinobacteria. Shotgun metagenomic sequencing further revealed functional traits of the rhizosphere microbiome in response to salt stress. This study enhances our understanding of mature tree-associated microbiomes and supports the theory that shaping the rhizosphere microbiome may be a strategy for saline-alkali-tolerant mature trees to resist salt stress. These findings provide insights into salt tolerance in mature trees and suggest potential applications, such as the development of bio-inoculants, for managing saline environments in agricultural and ecological contexts.
Keywords: mature pecan; microbiome; rhizosphere; salt stress; selectivity.
Copyright © 2025 Shi, Qin, Pu, Yang, Lim, Yang and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Exploring the Core Bacteria and Functional Traits in Pecan (Carya illinoinensis) Rhizosphere.Microbiol Spectr. 2023 Aug 17;11(4):e0011023. doi: 10.1128/spectrum.00110-23. Epub 2023 Jun 13. Microbiol Spectr. 2023. PMID: 37310220 Free PMC article.
-
Manipulating rhizosphere microorganisms to improve crop yield in saline-alkali soil: a study on soybean growth and development.Front Microbiol. 2023 Sep 20;14:1233351. doi: 10.3389/fmicb.2023.1233351. eCollection 2023. Front Microbiol. 2023. PMID: 37799597 Free PMC article.
-
Patterns in the Microbial Community of Salt-Tolerant Plants and the Functional Genes Associated with Salt Stress Alleviation.Microbiol Spectr. 2021 Oct 31;9(2):e0076721. doi: 10.1128/Spectrum.00767-21. Epub 2021 Oct 27. Microbiol Spectr. 2021. PMID: 34704793 Free PMC article.
-
Effects of saline-alkali stress on bacterial and fungal community diversity in Leymus chinensis rhizosphere soil.Environ Sci Pollut Res Int. 2022 Oct;29(46):70000-70013. doi: 10.1007/s11356-022-20270-6. Epub 2022 May 17. Environ Sci Pollut Res Int. 2022. PMID: 35579830
-
Microbially Mediated Plant Salt Tolerance and Microbiome-based Solutions for Saline Agriculture.Biotechnol Adv. 2016 Nov 15;34(7):1245-1259. doi: 10.1016/j.biotechadv.2016.08.005. Epub 2016 Aug 30. Biotechnol Adv. 2016. PMID: 27587331 Review.
References
LinkOut - more resources
Full Text Sources

