The genetics of low and high birthweight and their relationship with cardiometabolic disease
- PMID: 40210729
- PMCID: PMC12176956
- DOI: 10.1007/s00125-025-06420-8
The genetics of low and high birthweight and their relationship with cardiometabolic disease
Abstract
Aims/hypothesis: Low birthweight infants are at increased risk not only of mortality, but also of type 2 diabetes mellitus and CVD in later life. At the opposite end of the spectrum, high birthweight infants have increased risk of birth complications, such as shoulder dystocia, neonatal hypoglycaemia and obesity, and similarly increased risk of type 2 diabetes mellitus and CVD. However, previous genome-wide association studies (GWAS) of birthweight in the UK Biobank have primarily focused on individuals within the 'normal' range and have excluded individuals with high and low birthweight (<2.5 kg or >4.5 kg). The aim of this study was to investigate genetic variation associated within the tail ends of the birthweight distribution, to: (1) see whether the genetic factors operating in these regions were different from those that explained variation in birthweight within the normal range; (2) explore the genetic correlation between extremes of birthweight and cardiometabolic disease; and (3) investigate whether analysing the full distribution of birthweight values, including the extremes, improved the ability to detect genuine loci in GWAS.
Methods: We performed case-control GWAS analysis of low (<2.5 kg) and high (>4.5 kg) birthweight in the UK Biobank using REGENIE software (Nlow=20,947; Nhigh=12,715; Ncontrols=207,506) and conducted three continuous GWAS of birthweight, one including the full range of birthweights, one involving a truncated GWAS including only individuals with birthweights between 2.5 and 4.5 kg and a third GWAS that winsorised birthweight values <2.5 kg and >4.5 kg. Additionally, we performed bivariate linkage disequilibrium (LD) score regression to estimate the genetic correlation between low/normal/high birthweight and cardiometabolic traits.
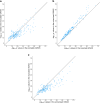
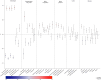
Results: Bivariate LD score regression analyses suggested that high birthweight had a mostly similar genetic aetiology to birthweight within the normal range (genetic correlation coefficient [rG]=0.91, 95% CI 0.83, 0.99), whereas there was more evidence for a separate set of genes underlying low birthweight (rG=-0.74, 95% CI 0.66, 0.82). Low birthweight was also significantly positively genetically correlated with most cardiometabolic traits and diseases we examined, whereas high birthweight was mostly positively genetically correlated with adiposity and anthropometric-related traits. The winsorisation strategy performed best in terms of locus detection, with the number of independent genome-wide significant associations (p<5×10-8) increasing from 120 genetic variants at 94 loci in the truncated GWAS to 270 genetic variants at 178 loci, including 27 variants at 25 loci that had not been identified in previous birthweight GWAS. This included a novel low-frequency missense variant in the ABCC8 gene, a gene known to be involved in congenital hyperinsulinism, neonatal diabetes mellitus and MODY, that was estimated to be responsible for a 170 g increase in birthweight amongst carriers.
Conclusions/interpretation: Our results underscore the importance of genetic factors in the genesis of the phenotypic correlation between birthweight and cardiometabolic traits and diseases.
Keywords: Birthweight; DOHaD; Developmental origins of health and disease; Diabetes; GWAS; Genome-wide association.
© 2025. The Author(s).
Conflict of interest statement
Acknowledgements: This research has been conducted using the UK Biobank resource (Reference 53641). We thank L. Yengo (University of Queensland, Australia), R. Freathy (University of Exeter, UK) and K. Birkeland (University of Oslo, Norway) for interesting discussions. Some of the data were presented as an abstract at the EPEC meeting in 2023. Data availability: Human genotype and phenotype data from the UK Biobank on which the results of this study were based were accessed with accession ID 53641. The genotype and phenotype data are available upon application to the UK Biobank ( http://www.ukbiobank.ac.uk/ ). Requirements for data access to the UK Biobank are described at https://www.ukbiobank.ac.uk/ . Funding: Open Access funding enabled and organized by CAUL and its Member Institutions. DME and this work are funded by an Australian National Health and Medical Research Council Investigator Grant (APP2017942) and Ideas Grant (GNT1183074). G-HM is the recipient of an Australian Research Council Discovery Early Career Award (Project number: DE220101226) funded by the Australian Government and supported by the Research Council of Norway (Project Grant: 325640). L-DH is funded by an Australian Research Council Discovery Early Career Researcher Award (DE240100014). NMW is funded by an Australian National Health and Medical Research Council Investigator Grant (APP2008723). Authors’ relationships and activities: The authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work. Contribution statement: G-HM made substantial contributions to the conception and design of the work, the acquisition, analysis and interpretation of data, as well as drafting the article. L-DH and NMW made substantial contributions to the interpretation of data and reviewing the article critically for important intellectual content. CBN made substantial contributions to the analysis of data and reviewing the article critically for important intellectual content. DME made substantial contributions to the conception and design of the work, the acquisition and interpretation of data, as well as drafting the article. All authors have approved the final version to be published. G-HM and DME are the guarantors of this work.
Figures
Similar articles
-
Meta-analysis of GWAS of over 16,000 individuals with autism spectrum disorder highlights a novel locus at 10q24.32 and a significant overlap with schizophrenia.Mol Autism. 2017 May 22;8:21. doi: 10.1186/s13229-017-0137-9. eCollection 2017. Mol Autism. 2017. PMID: 28540026 Free PMC article.
-
Structural variants linked to Alzheimer's disease and other common age-related clinical and neuropathologic traits.Genome Med. 2025 Mar 4;17(1):20. doi: 10.1186/s13073-025-01444-6. Genome Med. 2025. PMID: 40038788 Free PMC article.
-
Discovery and fine-mapping of adiposity loci using high density imputation of genome-wide association studies in individuals of African ancestry: African Ancestry Anthropometry Genetics Consortium.PLoS Genet. 2017 Apr 21;13(4):e1006719. doi: 10.1371/journal.pgen.1006719. eCollection 2017 Apr. PLoS Genet. 2017. PMID: 28430825 Free PMC article.
-
Prenatal administration of progestogens for preventing spontaneous preterm birth in women with a multiple pregnancy.Cochrane Database Syst Rev. 2019 Nov 20;2019(11):CD012024. doi: 10.1002/14651858.CD012024.pub3. Cochrane Database Syst Rev. 2019. PMID: 31745984 Free PMC article.
-
Pharmacological and electronic cigarette interventions for smoking cessation in adults: component network meta-analyses.Cochrane Database Syst Rev. 2023 Sep 12;9(9):CD015226. doi: 10.1002/14651858.CD015226.pub2. Cochrane Database Syst Rev. 2023. PMID: 37696529 Free PMC article.
References
-
- Hales CN, Barker DJ (1992) Type 2 (non-insulin-dependent) diabetes mellitus: the thrifty phenotype hypothesis. Diabetologia 35(7):595–601. 10.1007/BF00400248 - PubMed
-
- Roseboom TJ, van der Meulen JH, Ravelli AC, Osmond C, Barker DJ, Bleker OP (2001) Effects of prenatal exposure to the Dutch famine on adult disease in later life: an overview. Mol Cell Endocrinol 185(1–2):93–8 - PubMed
-
- Godfrey KM, Barker DJ (2000) Fetal nutrition and adult disease. Am J Clin Nutr 71(5):1344s–52s. 10.1093/ajcn/71.5.1344s - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

