Diurnal rhythms in durum wheat triggered by Rhopalosiphum padi (bird cherry-oat aphid)
- PMID: 40211135
- PMCID: PMC11984048
- DOI: 10.1186/s12870-025-06100-0
Diurnal rhythms in durum wheat triggered by Rhopalosiphum padi (bird cherry-oat aphid)
Abstract
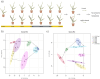
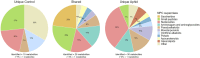
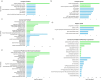
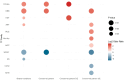
Wheat is a staple crop and one of the most widely consumed grains globally. Wheat yields can experience significant losses due to the damaging effects of herbivore infestation. However, little is known about the effect aphids have on the natural diurnal rhythms in plants. Our time-series transcriptomics and metabolomics study reveals intriguing molecular changes occurring in plant diurnal rhythmicity upon aphid infestation. Under control conditions, 15,366 out of the 66,559 genes in the tetraploid wheat cultivar Svevo, representing approximately 25% of the transcriptome, exhibited diurnal rhythmicity. Upon aphid infestation, 5,682 genes lost their rhythmicity, while 5,203 genes began to exhibit diurnal rhythmicity. The aphid-induced rhythmic genes were enriched in GO terms associated with plant defense, such as protein phosphorylation and cellular response to ABA and were enriched with motifs of the WRKY transcription factor families. In contrast, the genes that lost rhythmicity due to aphid infestation were enriched with motifs of the TCP and ERF transcription factor families. While the core circadian clock genes maintain their rhythmicity during infestation, we observed that approximately 60% of rhythmic genes experience disruptions in their rhythms during aphid infestation. These changes can influence both the plant's growth and development processes as well as defense responses. Furthermore, analysis of rhythmic metabolite composition revealed that several monoterpenoids gained rhythmic activity under infestation, while saccharides retained their rhythmic patterns. Our findings highlight the ability of insect infestation to disrupt the natural diurnal cycles in plants, expanding our knowledge of the complex interactions between plants and insects.
Keywords: Rhopalosiphum padi; Aphid; Diurnal; Rhythmicity; Transcription factor; Transcriptomics; Untargeted metabolomics; WRKY; Wheat.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures

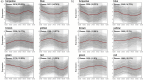

References
-
- Nozue K, Maloof JN. Diurnal regulation of plant growth. Plant Cell Environ. 2006;29. - PubMed
-
- Erenstein O, Jaleta M, Mottaleb KA, Sonder K, Donovan J, Braun H-J. Global trends in Wheat Production, Consumption and Trade. In: Reynolds MP, Braun H-J, editors. Wheat improvement: Food Security in a changing climate. Cham: Springer International Publishing; 2022. pp. 47–66.
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

