Reduction of alternative polarization of macrophages by short-term activated hepatic stellate cell-derived small extracellular vesicles
- PMID: 40211350
- PMCID: PMC11983935
- DOI: 10.1186/s13046-025-03380-0
Reduction of alternative polarization of macrophages by short-term activated hepatic stellate cell-derived small extracellular vesicles
Abstract
Background: Activated hepatic stellate cells (HSCs) induce alternative (M2) polarization of macrophages and contribute to the progression of fibrosis and hepatocellular carcinoma (HCC). However, the effects of small extracellular vesicles released by HSCs (HSC-sEVs) during activation remain largely unknown.
Methods: The aim of this study was to investigate the role of extracellular vesicles released by HSCs (HSC-sEVs) at different stages of activation in macrophage polarization. The effects of sEVs from short-term activated and long-term activated HSCs on liver macrophages was studied. Small RNA sequencing analyses were performed to obtain differential miRNAs transported by the short-term and long-term activated HSC- sEVs. The in vivo effects of short-term activated HSC-sEV-specific miRNA on liver macrophage and liver fibrosis were confirmed in a CCl4-induced liver injury mouse model. To study the tumor suppressive effects of the macrophages educated by short-term activated HSC-sEV-specific miRNA, human hepatoma cells were mixed and subcutaneously cotransplanted with miR-99a-5p mimic-pretreated macrophages.
Results: We found that consistent with activated HSCs, long-term activated HSC-sEVs (14dHSC-sEVs) induce bone marrow-derived monocytes (MOs) toward an M2 phenotype, but short-term activated HSC-sEVs (3dHSC-sEVs) induce the resident macrophages (Kupffer cells, KCs) toward a classically activated (M1) phenotype. We identified five 3dHSC-sEV-specific miRNAs, including miR-99a-5p. In vitro and in vivo experiments support that miR-99a-5p negatively regulates alternative polarization of macrophages, decreases collagen deposition in chronic liver injury model, and suppresses the progression of hepatoma in a xenograft model partially by targeting CD93.
Conclusion: Collectively, our work reveals an unexpected proinflammatory role of 3dHSC-sEVs, preliminarily explores the underlying mechanism, and evaluates the therapeutic potential of 3dHSC-sEV-specific miR-99a-5p for liver fibrosis and tumorigenesis.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures





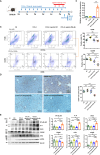


References
-
- Kim JW, Kim YJ. The evidence-based multifaceted roles of hepatic stellate cells in liver diseases: A concise review. Life Sci 2024;344:122547. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

