The Atypical Pectin Methylesterase Family Member PME31 Promotes Seedling Lipid Droplet Utilization
- PMID: 40212536
- PMCID: PMC11982519
- DOI: 10.1002/pld3.70054
The Atypical Pectin Methylesterase Family Member PME31 Promotes Seedling Lipid Droplet Utilization
Abstract
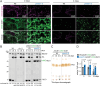
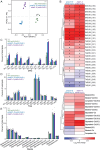
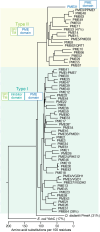
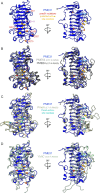
In plants, the primary form of energy stored in seed lipid droplets, triacylglycerol (TAG), is catabolized during germination to support pre-photosynthetic growth. Although this process is essential for seedling development, it is incompletely understood. In a screen for Arabidopsis thaliana mutants displaying delayed degradation of the lipid droplet coat protein oleosin, five independent mutations in PECTIN METHYLESTERASE31 (PME31) were recovered. In addition to delayed oleosin degradation, pme31 mutant seedlings exhibited sustained lipid droplets and elevated levels of several TAG and diacylglycerol species. Although structural prediction classified PME31 as a pectinesterase, this structural family also includes a putative E. coli lipase, YbhC. Moreover, PME31 lacks an N-terminal signal peptide that would target it to the cell wall, where pectin resides. We found that a fluorescent PME31 reporter was cytosolic and partially associated with peroxisomes, the site of fatty acid catabolism, during lipid mobilization. Our findings suggest that, in contrast to canonical PMEs, which modify cell wall pectin, PME31 functions at peroxisomes to directly or indirectly promote lipid mobilization.
Keywords: Arabidopsis germination; lipid droplet; lipid mobilization; oleosin; pectin methylesterase; peroxisome.
© 2025 The Author(s). Plant Direct published by American Society of Plant Biologists and the Society for Experimental Biology and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
The ubiquitin-protein ligase MIEL1 localizes to peroxisomes to promote seedling oleosin degradation and lipid droplet mobilization.Proc Natl Acad Sci U S A. 2023 Jul 18;120(29):e2304870120. doi: 10.1073/pnas.2304870120. Epub 2023 Jul 6. Proc Natl Acad Sci U S A. 2023. PMID: 37410814 Free PMC article.
-
Pectin methylesterase 31 is transcriptionally repressed by ABI5 to negatively regulate ABA-mediated inhibition of seed germination.Front Plant Sci. 2024 Feb 2;15:1336689. doi: 10.3389/fpls.2024.1336689. eCollection 2024. Front Plant Sci. 2024. PMID: 38371403 Free PMC article.
-
Pectin methylesterase31 positively regulates salt stress tolerance in Arabidopsis.Biochem Biophys Res Commun. 2018 Feb 5;496(2):497-501. doi: 10.1016/j.bbrc.2018.01.025. Epub 2018 Jan 4. Biochem Biophys Res Commun. 2018. PMID: 29307824
-
Exploiting lipid droplet metabolic pathway to foster lipid production: oleosin in focus.Plant Cell Rep. 2024 Dec 26;44(1):12. doi: 10.1007/s00299-024-03390-w. Plant Cell Rep. 2024. PMID: 39724216 Review.
-
Biogenesis and Lipase-Mediated Mobilization of Lipid Droplets in Plants.Plants (Basel). 2022 May 5;11(9):1243. doi: 10.3390/plants11091243. Plants (Basel). 2022. PMID: 35567244 Free PMC article. Review.
References
-
- Adham, A. R. , Zolman B. K., Millius A., and Bartel B.. 2005. “Mutations in Arabidopsis Acyl‐CoA Oxidase Genes Reveal Distinct and Overlapping Roles in β‐Oxidation.” Plant Journal 41: 859–874. - PubMed
-
- Anderson, C. T. 2016. “We Be Jammin': An Update on Pectin Biosynthesis, Trafficking and Dynamics.” Journal of Experimental Botany 67: 495–502. - PubMed
-
- Clough, S. J. , and Bent A. F.. 1998. “Floral Dip: A Simplified Method for Agrobacterium‐Mediated Transformation of Arabidopsis thaliana .” Plant Journal 16: 735–743. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

