Decoding the impact of interspecies interactions on biofilm matrix components
- PMID: 40212915
- PMCID: PMC11985002
- DOI: 10.1016/j.bioflm.2025.100271
Decoding the impact of interspecies interactions on biofilm matrix components
Abstract
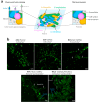
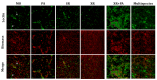
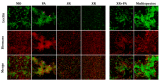
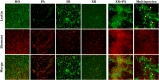
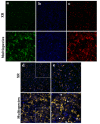
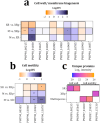
Multispecies biofilms are complex communities where extracellular polymeric substances (EPS) shape structure, adaptability, and functionality. However, characterizing the components of EPS, particularly glycans and proteins, remains a challenge due to the diverse bacterial species present and their interactions within the matrix. This study examined how interactions between different species affect EPS component composition and spatial organization. We analyzed a consortium of four bacterial soil isolates that have previously demonstrated various intrinsic properties in biofilm communities: Microbacterium oxydans, Paenibacillus amylolyticus, Stenotrophomonas rhizophila, and Xanthomonas retroflexus. We used fluorescence lectin binding analysis to identify specific glycan components and meta-proteomics to characterize matrix proteins in mono- and multispecies biofilms. Our results revealed diverse glycan structures and composition, including fucose and different amino sugar-containing polymers, with substantial differences between monospecies and multispecies biofilms. In isolation, M. oxydans produced galactose/N-Acetylgalactosamine network-like structures and influenced the matrix composition in multispecies biofilms. Proteomic analysis revealed presence of flagellin proteins in X. retroflexus and P. amylolyticus, particularly in multispecies biofilms. Additionally, surface-layer proteins and a unique peroxidase were identified in P. amylolyticus multispecies biofilms, indicating enhanced oxidative stress resistance and structural stability under these conditions. This study highlights the crucial role of interspecies interactions in shaping the biofilm matrix, as well as the production of glycans and proteins, thereby enhancing our understanding of biofilm complexity.
Keywords: Extracellular polymeric substances; Glycans; Interspecies interactions; Multispecies biofilms.
© 2025 The Authors.
Conflict of interest statement
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Mette Burmolle reports financial support was provided by European Research Council. Mette Burmølle reports financial support was provided by Office of Naval Research. Henriette Lyng Røder reports financial support was provided by Villum Foundation. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

References
-
- Flemming H.C., Wingender J., Szewzyk U., Steinberg P., Rice S.A., Kjelleberg S. Biofilms: an emergent form of bacterial life. Nat Rev Microbiol. 2016;14:563–575. Nature Publishing Group. - PubMed
-
- Flemming H.C., van Hullebusch E.D., Neu T.R., Nielsen P.H., Seviour T., Stoodley P., et al. The biofilm matrix: multitasking in a shared space. Nat Rev Microbiol. 2022;21:70–86. 2022 212. - PubMed
-
- Branda S.S., Vik Å., Friedman L., Kolter R. Biofilms: the matrix revisited. Trends Microbiol. 2005;13:20–26. - PubMed
-
- Flemming H.C., Wingender J. The biofilm matrix. Nat Rev Microbiol. 2010;8:623–633. - PubMed
-
- Karygianni L., Ren Z., Koo H., Thurnheer T. Biofilm matrixome: extracellular components in structured microbial communities. Trends Microbiol. 2020;28:668–681. - PubMed
LinkOut - more resources
Full Text Sources

