Astrocytic RNA editing regulates the host immune response to alpha-synuclein
- PMID: 40215316
- PMCID: PMC11988446
- DOI: 10.1126/sciadv.adp8504
Astrocytic RNA editing regulates the host immune response to alpha-synuclein
Abstract
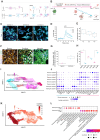
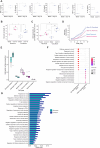
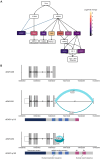
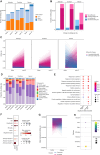
RNA editing is a posttranscriptional mechanism that targets changes in RNA transcripts to modulate innate immune responses. We report the role of astrocyte-specific, ADAR1-mediated RNA editing in neuroinflammation in Parkinson's disease (PD). We generated human induced pluripotent stem cell-derived astrocytes, neurons and cocultures and exposed them to small soluble alpha-synuclein aggregates. Oligomeric alpha-synuclein triggered an inflammatory glial state associated with Toll-like receptor activation, viral responses, and cytokine secretion. This reactive state resulted in loss of neurosupportive functions and the induction of neuronal toxicity. Notably, interferon response pathways were activated leading to up-regulation and isoform switching of the RNA deaminase enzyme, ADAR1. ADAR1 mediates A-to-I RNA editing, and increases in RNA editing were observed in inflammatory pathways in cells, as well as in postmortem human PD brain. Aberrant, or dysregulated, ADAR1 responses and RNA editing may lead to sustained inflammatory reactive states in astrocytes triggered by alpha-synuclein aggregation, and this may drive the neuroinflammatory cascade in Parkinson's.
Figures

References
-
- Braak H., Del Tredici K., Rüb U., de Vos R. A. I., Jansen Steur E. N. H., Braak E., Staging of brain pathology related to sporadic Parkinson’s disease. Neurobiol. Aging 24, 197–211 (2003). - PubMed
-
- Damier P., Hirsch E. C., Zhang P., Agid Y., Javoy-Agid F., Glutathione peroxidase, glial cells and Parkinson’s disease. Neuroscience 52, 1–6 (1993). - PubMed
-
- Knott C., Wilkin G. P., Stern G., Astrocytes and microglia in the substantia nigra and caudate-putamen in Parkinson’s disease. Parkinsonism Relat. Disord. 5, 115–122 (1999). - PubMed
-
- Miklossy J., Doudet D. D., Schwab C., Yu S., McGeer E. G., McGeer P. L., Role of ICAM-1 in persisting inflammation in Parkinson disease and MPTP monkeys. Exp. Neurol. 197, 275–283 (2006). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

