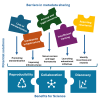
Perceptual and technical barriers in sharing and formatting metadata accompanying omics studies
- PMID: 40215974
- PMCID: PMC12143318
- DOI: 10.1016/j.xgen.2025.100845
Perceptual and technical barriers in sharing and formatting metadata accompanying omics studies
Abstract
Metadata, or "data about data," is essential for organizing, understanding, and managing large-scale omics datasets. It enhances data discovery, integration, and interpretation, enabling reproducibility, reusability, and secondary analysis. However, metadata sharing remains hindered by perceptual and technical barriers, including the lack of uniform standards, privacy concerns, study design limitations, insufficient incentives, inadequate infrastructure, and a shortage of trained personnel. These challenges compromise data reliability and obstruct integrative meta-analyses. Addressing these issues requires standardization, education, stronger roles for journals and funding agencies, and improved incentives and infrastructure. Looking ahead, emerging technologies such as artificial intelligence and machine learning may offer promising solutions to automate metadata processes, increasing accuracy and scalability. Fostering a collaborative culture of metadata sharing will maximize the value of omics data, accelerating innovation and scientific discovery.
Keywords: barriers in metadata sharing practices; data; metadata; metadata completeness.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Similar articles
-
Proposal for Using AI to Assess Clinical Data Integrity and Generate Metadata: Algorithm Development and Validation.JMIR Med Inform. 2025 Jun 30;13:e60204. doi: 10.2196/60204. JMIR Med Inform. 2025. PMID: 40587839 Free PMC article.
-
A Case for Accelerating Standards to Achieve the FAIR Principles of Environmental Health Research Experimental Data.Environ Health Perspect. 2023 Jun;131(6):65001. doi: 10.1289/EHP11484. Epub 2023 Jun 23. Environ Health Perspect. 2023. PMID: 37352010 Free PMC article.
-
The Use of AI for Phenotype-Genotype Mapping.Methods Mol Biol. 2025;2952:369-410. doi: 10.1007/978-1-0716-4690-8_21. Methods Mol Biol. 2025. PMID: 40553344
-
A Systematic Review and Bibliometric Analysis of Applications of Artificial Intelligence and Machine Learning in Vascular Surgery.Ann Vasc Surg. 2022 Sep;85:395-405. doi: 10.1016/j.avsg.2022.03.019. Epub 2022 Mar 24. Ann Vasc Surg. 2022. PMID: 35339595
-
Navigating Challenges in the Integration of Artificial Intelligence in Nursing Practice: An Integrative Literature Review.Int Nurs Rev. 2025 Sep;72(3):e70077. doi: 10.1111/inr.70077. Int Nurs Rev. 2025. PMID: 40685999 Review.
Cited by
-
Assessing the safety of microbiome perturbations.Microb Genom. 2025 May;11(5):001405. doi: 10.1099/mgen.0.001405. Microb Genom. 2025. PMID: 40371892 Free PMC article. Review.
-
Harnessing Multi-Omics and Predictive Modeling for Climate-Resilient Crop Breeding: From Genomes to Fields.Genes (Basel). 2025 Jul 10;16(7):809. doi: 10.3390/genes16070809. Genes (Basel). 2025. PMID: 40725465 Free PMC article. Review.
-
The systematic assessment of completeness of public metadata accompanying omics studies in the Gene Expression Omnibus.bioRxiv [Preprint]. 2025 Jul 7:2021.11.22.469640. doi: 10.1101/2021.11.22.469640. bioRxiv. 2025. PMID: 40672350 Free PMC article. Preprint.
-
UShER-TB: Scalable, Comprehensive, Accessible Phylogenomic Analysis of Mycobacterium tuberculosis.medRxiv [Preprint]. 2025 Jul 23:2025.07.22.25331806. doi: 10.1101/2025.07.22.25331806. medRxiv. 2025. PMID: 40778146 Free PMC article. Preprint.
-
A guide to developing harmonized research workflows in a team science context.Exp Neurol. 2025 Oct;392:115333. doi: 10.1016/j.expneurol.2025.115333. Epub 2025 Jun 5. Exp Neurol. 2025. PMID: 40482901 Free PMC article. Review.
References
-
- Huang Y.-N., Patel N.A., Mehta J.H., Ginjala S., Brodin P., Gray C.M., Patel Y.M., Cowell L.G., Burkhardt A.M., Mangul S. Data Availability of Open T-Cell Receptor Repertoire Data, a Systematic Assessment. Front. Syst. Biol. 2022;2 doi: 10.3389/fsysb.2022.918792. - DOI
-
- Clough E., Barrett T. In: Statistical Genomics: Methods and Protocols. Mathé E., Davis S., editors. Springer; 2016. The Gene Expression Omnibus Database; pp. 93–110. - DOI
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources