Integrated single cell and bulk RNA sequencing analyses reveal the impact of tryptophan metabolism on prognosis and immunotherapy in colon cancer
- PMID: 40216815
- PMCID: PMC11992224
- DOI: 10.1038/s41598-025-85893-4
Integrated single cell and bulk RNA sequencing analyses reveal the impact of tryptophan metabolism on prognosis and immunotherapy in colon cancer
Abstract
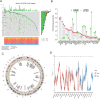
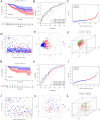
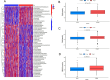
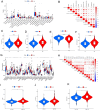
Tryptophan metabolism is intricately associated with the progression of colon cancer. This research endeavored to meticulously analyze tryptophan metabolic characteristics in colon cancer and forecast immunotherapy responses. This study analyzed colon cancer samples from a training cohort of 473 tumors and 41 normal tissues from TCGA, with validation in 902 cancer patients across multiple GEO datasets. Patients were stratified into subtypes through consistent clustering, and a tryptophan metabolic risk score model was constructed using the random forest algorithm. Based on these risk scores, patients were delineated into high and low-risk groups, and their clinicopathologic characteristics, immune cell infiltration, immune checkpoint expression, and signaling pathway disparities were examined. The Oncopredict algorithm facilitated the identification of sensitive chemotherapeutic agents, while the immune escape score was employed to evaluate the immunotherapy response across risk groups. Transcriptomic sequencing findings were corroborated by single-cell sequencing from Shanghai Ruijin Hospital. Two distinct subtypes of colon cancer patients emerged, exhibiting significant prognostic and immune cell infiltration differences. The high-risk group demonstrated a poorer prognosis (p < 0.001), advanced clinical stage (p < 0.001), and elevated immunosuppressive cell expression (p < 0.05). Additionally, three chemotherapeutic drugs showed efficacy in the high-risk cohort, displaying a heightened immune escape potential (p < 0.05) and diminished response to immunotherapy. Single-cell sequencing validated the overexpression of tryptophan-related genes in epithelial cells. In conclusion, tryptophan metabolism significantly influences the colon cancer immune microenvironment, with high-risk patients experiencing adverse prognoses and potentially reduced efficacy of immunotherapy.
Keywords: Colon cancer; Immunotherapy; Prognosis; Risk score; Tryptophan metabolism.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures



Similar articles
-
Identification of a metabolic-immune signature associated with prognosis in colon cancer and exploration of potential predictive efficacy of immunotherapy response.Clin Exp Med. 2025 Jan 24;25(1):46. doi: 10.1007/s10238-025-01566-6. Clin Exp Med. 2025. PMID: 39853414 Free PMC article.
-
Integrated analysis of bulk and single-cell RNA sequencing reveals the impact of nicotinamide and tryptophan metabolism on glioma prognosis and immunotherapy sensitivity.BMC Neurol. 2024 Oct 28;24(1):419. doi: 10.1186/s12883-024-03924-5. BMC Neurol. 2024. PMID: 39468708 Free PMC article.
-
Development and validation of a cancer-associated fibroblast gene signature-based model for predicting immunotherapy response in colon cancer.Sci Rep. 2025 May 13;15(1):16550. doi: 10.1038/s41598-025-01185-x. Sci Rep. 2025. PMID: 40360558 Free PMC article.
-
Identification and validation of tryptophan-related gene signatures to predict prognosis and immunotherapy response in lung adenocarcinoma reveals a critical role for PTTG1.Front Immunol. 2024 Jul 31;15:1386427. doi: 10.3389/fimmu.2024.1386427. eCollection 2024. Front Immunol. 2024. PMID: 39144144 Free PMC article.
-
Constructing and validating a risk model based on neutrophil-related genes for evaluating prognosis and guiding immunotherapy in colon cancer.J Gene Med. 2024 Apr;26(4):e3684. doi: 10.1002/jgm.3684. J Gene Med. 2024. PMID: 38618694 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

