Development of an automated plaque-counting program for the quantification of the Chikungunya virus
- PMID: 40217093
- PMCID: PMC11992114
- DOI: 10.1038/s41598-025-97590-3
Development of an automated plaque-counting program for the quantification of the Chikungunya virus
Abstract
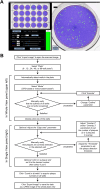
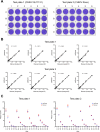
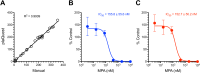
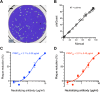
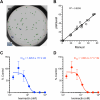
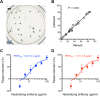
Chikungunya virus (CHIKV) induces a massive cytopathic effect (CPE) on various cell types. Therefore, the plaque assay, a CPE-based virus titration method, remains the gold standard for quantifying the infectious units of CHIKV. However, manual plaque counting is often a labor-intensive task, especially in experiments involving multiple samples. In this study, we developed plaQuest, a stand-alone plaque-counting software running on a Windows operating system, for rapid and reliable quantification of CHIKV plaques in a 24-well plate. Our evaluation experiments using the conventional CPE-based plaque assay showed that the CHIKV plaque counts provided by plaQuest strongly correlated with the plaque counts manually determined by four analysts. In addition, the CHIKV inhibition curve of mycophenolic acid (MPA) determined by plaQuest was identical to that determined by manual counting, resulting in a similar 50% inhibitory concentration of MPA. Furthermore, the automated plaque counting by plaQuest was applicable to the evaluation of inhibitors against other RNA viruses using the CPE-based and immunostain-based plaque assay, which is an alternative titration assay for non- (or less) cytopathic viruses. Thus, our study demonstrates that plaQuest is an effective option for quantifying infectious virus titers, reducing the workload of the plaque assay.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures
References
-
- Condit, R. C. Principles of virology. In Fields Virology, 5th edn (eds Knipe, D. M. & Howley, P. M.) Vol. I 25–57 (Lippincott Williams & Wilkins, 2007).
-
- The Science of Virology. In Principles of Virology: Molecular Biology, Pathogenesis, and Control 29–31 (eds Flint, S. J., Enquist, L. W., Krug, R. M., Racaniello, V. R. & Skalka) (ASM, 2000).
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

