The RMaP challenge of predicting RNA modifications by nanopore sequencing
- PMID: 40221591
- PMCID: PMC11993749
- DOI: 10.1038/s42004-025-01507-0
The RMaP challenge of predicting RNA modifications by nanopore sequencing
Abstract
The field of epitranscriptomics is undergoing a technology-driven revolution. During past decades, RNA modifications like N6-methyladenosine (m6A), pseudouridine (ψ), and 5-methylcytosine (m5C) became acknowledged for playing critical roles in cellular processes. Direct RNA sequencing by Oxford Nanopore Technologies (ONT) enabled the detection of modifications in native RNA, by detecting noncanonical RNA nucleosides properties in raw data. Consequently, the field's cutting edge has a heavy component in computer science, opening new avenues of cooperation across the community, as exchanging data is as impactful as exchanging samples. Therefore, we seize the occasion to bring scientists together within the RNA Modification and Processing (RMaP) challenge to advance solutions for RNA modification detection and discuss ideas, problems and approaches. We show several computational methods to detect the most researched mRNA modifications (m6A, ψ, and m5C). Results demonstrate that a low prediction error and a high prediction accuracy can be achieved on these modifications across different approaches and algorithms. The RMaP challenge marks a substantial step towards improving algorithms' comparability, reliability, and consistency in RNA modification prediction. It points out the deficits in this young field that need to be addressed in further challenges.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests. Mark Helm is a consultant for Moderna Inc.
Figures

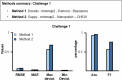



References
-
- Delaunay, S., Helm, M. & Frye, M. RNA modifications in physiology and disease: towards clinical applications. Nat. Rev. Genet.25, 104–122 (2024). - PubMed
-
- Lucas, M. C. & Novoa, E. M. Long-read sequencing in the era of epigenomics and epitranscriptomics. Nat. Methods20, 25–29 (2023). - PubMed
-
- Alfonzo, J. D. et al. A call for direct sequencing of full-length RNAs to identify all modifications. Nat. Genet.53, 1113–1116 (2021). - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

