circHIPK2 promotes malignant progression of laryngeal squamous cell carcinoma through the miR-889-3p/MCTS1/IL-6 axis
- PMID: 40222337
- PMCID: PMC12018564
- DOI: 10.1016/j.tranon.2025.102390
circHIPK2 promotes malignant progression of laryngeal squamous cell carcinoma through the miR-889-3p/MCTS1/IL-6 axis
Abstract
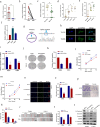
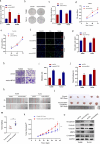
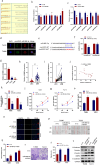
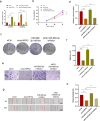
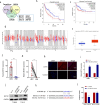
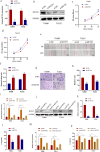
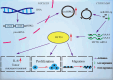
Laryngeal squamous cell carcinoma (LSCC) is a common malignant tumor of the head and neck with a poor prognosis. The role of circRNAs in LSCC remains largely unknown. In this study, quantitative real-time PCR (qRT-PCR), Sanger sequencing and fluorescence in situ hybridization were undertaken to detect the expression, localization, and clinical significance of circHIPK2 in LSCC tissues and TU686 and TU212 cells. The functions of circHIPK2 in LSCC were explored through proliferation analysis, EdU staining, colony formation assay, wound healing assay, and Transwell assay. The regulatory mechanisms underpinning circHIPK2, miR-889-3p, and MCTS1 were investigated using luciferase assay, Western blotting, and qRT-PCR. We found that LSCC tissues and cells demonstrated high expression of circHIPK2 that was closely associated with the malignant progression and poor prognosis of LSCC. Knockdown of circHIPK2 inhibited the proliferation and migration of LSCC cells in vitro. Mechanistic studies showed that circHIPK2 competitively bound to miR-889-3p, elevated MCTS1 level, promoted IL-6 secretion, and ultimately accelerated the malignant progression of LSCC. In conclusion, an axis involving circHIPK2, miR-889-3p, MCTS1 and IL-6 regulates the malignant progression of LSCC. circHIPK2 expression may serve as a novel diagnostic and prognostic biomarker for LSCC.
Keywords: IL-6; Laryngeal squamous cell carcinoma; MCTS1; Non-coding RNA; Tumor microenvironment; circHIPK2; miR-889–3p.
Copyright © 2025. Published by Elsevier Inc.
Conflict of interest statement
Declaration of competing interest The authors declare that they have no competing interests.
Figures
Similar articles
-
Circular RNA circCORO1C promotes laryngeal squamous cell carcinoma progression by modulating the let-7c-5p/PBX3 axis.Mol Cancer. 2020 Jun 2;19(1):99. doi: 10.1186/s12943-020-01215-4. Mol Cancer. 2020. PMID: 32487167 Free PMC article.
-
PTPN12 down-regulated by miR-146b-3p gene affects the malignant progression of laryngeal squamous cell carcinoma.Open Med (Wars). 2023 Jun 9;18(1):20230727. doi: 10.1515/med-2023-0727. eCollection 2023. Open Med (Wars). 2023. PMID: 37333450 Free PMC article.
-
Circ_0023028 contributes to the progression of laryngeal squamous cell carcinoma by upregulating LASP1 through miR-486-3p.Mol Cell Biochem. 2021 Aug;476(8):2951-2961. doi: 10.1007/s11010-021-04129-x. Epub 2021 Mar 23. Mol Cell Biochem. 2021. PMID: 33755879
-
Hsa_circ_0042823 accelerates cancer progression via miR-877-5p/FOXM1 axis in laryngeal squamous cell carcinoma.Ann Med. 2021 Dec;53(1):960-970. doi: 10.1080/07853890.2021.1934725. Ann Med. 2021. PMID: 34124974 Free PMC article.
-
Dysbiosis Patterns in Glottic and Laryngeal Cancers: A Systematic Review of Microbiome Alterations.J Voice. 2025 Mar 10:S0892-1997(25)00083-9. doi: 10.1016/j.jvoice.2025.02.036. Online ahead of print. J Voice. 2025. PMID: 40069023 Review.
References
-
- Lv Y., Wang Y., Zhang Z. Potentials of lncRNA-miRNA-mRNA networks as biomarkers for laryngeal squamous cell carcinoma. Hum. Cell. 2023;36:76–97. - PubMed
-
- Santos A., Santos I.C., Dos Reis P.F., Rodrigues V.D., Peres W.A.F. Impact of Nutritional Status on Survival in Head and Neck Cancer Patients After Total Laryngectomy. Nutr. Cancer. 2022;74:1252–1260. - PubMed
-
- Baird B.J., Sung C.K., Beadle B.M., Divi V. Treatment of early-stage laryngeal cancer: a comparison of treatment options. Oral Oncol. 2018;87:8–16. - PubMed
LinkOut - more resources
Full Text Sources

