A species-level identification pipeline for human gut microbiota based on the V3-V4 regions of 16S rRNA
- PMID: 40226098
- PMCID: PMC11985812
- DOI: 10.3389/fmicb.2025.1553124
A species-level identification pipeline for human gut microbiota based on the V3-V4 regions of 16S rRNA
Abstract
16S rRNA gene sequencing is pivotal for identifying bacterial species in microbiome studies, especially using the V3-V4 hypervariable regions. A fixed 98.5% similarity threshold is often applied for species-level identification, but this approach can cause misclassification due to varying thresholds among species. To address this, our study integrated data from SILVA, NCBI, and LPSN databases, extracting V3-V4 region sequences and supplementing them with 16S rRNA sequences from 1,082 human gut samples. This resulted in a non-redundant amplicon sequence variants (ASVs) database specific to the V3-V4 regions (positions 341-806). Utilizing this database, we identified flexible classification thresholds for 674 families, 3,661 genera, and 15,735 species, finding clear thresholds for 87.09% of families and 98.38% of genera. For the 896 most common human gut species, we established precise taxonomic thresholds. To leverage these findings, we developed the asvtax pipeline, which applies flexible thresholds for more accurate taxonomic classification, notably improving the identification of new ASVs. The asvtax pipeline not only enhances the precision of species-level classification but also provides a robust framework for analyzing complex microbial communities, facilitating more reliable ecological and functional interpretations in microbiome research.
Keywords: 16S rRNA; database abbreviations; microbiota; species-level identification; taxonomic thresholds.
Copyright © 2025 Wang, Yuan, Chen, Yang, Pu, Lin, Dong, Zhang, Yuan, Zheng, Sun and Xu.
Conflict of interest statement
WL was employed by Uniteomics Tianjin Biotechnology Co., Ltd. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

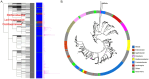

References
-
- Chiarello M., McCauley M., Villéger S., Jackson C. R. (2022). Ranking the biases: the choice of OTUs vs. ASVs in 16S rRNA amplicon data analysis has stronger effects on diversity measures than rarefaction and OTU identity threshold. PLoS One 17:e0264443. doi: 10.1371/journal.pone.0264443, PMID: - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

