HBO1 determines epithelial-mesenchymal transition and promotes immunotherapy resistance in ovarian cancer cells
- PMID: 40227530
- PMCID: PMC12238117
- DOI: 10.1007/s13402-025-01055-8
HBO1 determines epithelial-mesenchymal transition and promotes immunotherapy resistance in ovarian cancer cells
Abstract
Purpose: Epithelial-mesenchymal transition (EMT) plays critical roles in tumor progress and treatment resistance of ovarian cancer (OC), resulting in the most deadly gynecological cancer in women. However, the cell-intrinsic mechanism underlying EMT in OC remains less illuminated.
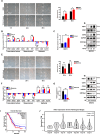
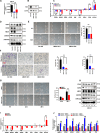
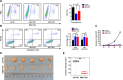
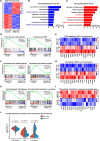
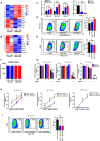
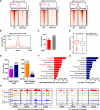
Method: SKOV3, the OC cell line, was treated with TGF-β to induce EMT or with SB431542, an inhibitor of the TGF-β signaling pathway, to reduce migration. The function of HBO1 in EMT was confirmed by knock-down or overexpression of HBO1 in SKOV3 cells. The role of HBO1 in cell proliferation and apoptosis of SKOV3 cells was analyzed by flow cytometry. The whole-genome transcriptome was used to compare significantly different genes in control and HBO1-KD SKOV3 cells. T-cell cytotoxicity assays were measured by an IVIS spectrum. The chromatin binding of HBO1 was investigated using CUT&Tag-seq.
Results: Here, we show that HBO1, a MYST histone acetyltransferase (HAT), is a cell-intrinsic determinant for EMT in OC cells. HBO1 is greatly elevated during TGF-β-triggered EMT in SKOV3 OC cells as well as in later stages of clinical OC samples. HBO1 Knock-down (KD) in SKOV3 cells blocks TGF-β-triggered EMT, migration, invasion and tumor formation in vivo. Interestingly, HBO1 KD in SKOV3 cells suppresses their resistance to CAR-T cells. Mechanistically, HBO1 co-binds the gene sets responsible for EMT with SMAD4 and orchestrates a gene regulatory network critical for tumor progression in SKOV3 cells.
Conclusion: HBO1 plays an essential onco-factor to drive EMT and promote the immunotherapy resistance in ovarian cancer cells. Together, we reveal a critical role of HBO1 mediated epigenetic mechanism in OC progression, providing an insight into designing new therapy strategies.
Keywords: Epithelial-mesenchymal transition; HBO1; Oncogene; Ovarian cancer; TGF-β signaling.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethical approval consent to publish: All animal experiments were conducted in accordance with an approved protocol by the Institutional Animal Care and Use Committee (IACUC) at the Guangzhou Institutes of Biomedicine and Health, Chinese academy of science. Competing interests: The authors declare no competing interests.
Figures

Similar articles
-
MYB/AKT3 axis is a key driver of ovarian cancer growth, aggressiveness, and chemoresistance.J Ovarian Res. 2025 Aug 11;18(1):179. doi: 10.1186/s13048-025-01761-9. J Ovarian Res. 2025. PMID: 40790586 Free PMC article.
-
THEMIS2 contributes to ovarian cancer metastasis via DOCK4-mediated activation of Rap1 signaling.Cell Oncol (Dordr). 2025 Aug;48(4):961-978. doi: 10.1007/s13402-025-01057-6. Epub 2025 Apr 14. Cell Oncol (Dordr). 2025. PMID: 40227532 Free PMC article.
-
Human chorionic gonadotropin β regulates epithelial-mesenchymal transition and metastasis in human ovarian cancer.Oncol Rep. 2017 Sep;38(3):1464-1472. doi: 10.3892/or.2017.5818. Epub 2017 Jul 14. Oncol Rep. 2017. PMID: 28713970 Free PMC article.
-
Impact of residual disease as a prognostic factor for survival in women with advanced epithelial ovarian cancer after primary surgery.Cochrane Database Syst Rev. 2022 Sep 26;9(9):CD015048. doi: 10.1002/14651858.CD015048.pub2. Cochrane Database Syst Rev. 2022. PMID: 36161421 Free PMC article.
-
Intraoperative frozen section analysis for the diagnosis of early stage ovarian cancer in suspicious pelvic masses.Cochrane Database Syst Rev. 2016 Mar 1;3(3):CD010360. doi: 10.1002/14651858.CD010360.pub2. Cochrane Database Syst Rev. 2016. PMID: 26930463 Free PMC article.
References
-
- L.E. Kandalaft, D. Dangaj Laniti, G. Coukos, Nat. Rev. Cancer 22, 640–656 (2022). 10.1038/s41568-022-00503-z - PubMed
-
- S. Vaughan, J.I. Coward, R.C. Bast Jr, A. Berchuck, J.S. Berek, J.D. Brenton, G. Coukos, C.C. Crum, R. Drapkin, D. Etemadmoghadam, M. Friedlander, H. Gabra, S.B. Kaye, C.J. Lord, E. Lengyel, D.A. Levine, I.A. McNeish, U. Menon, G.B. Mills, K.P. Nephew, A.M. Oza, A.K. Sood, E.A. Stronach, H. Walczak, D.D. Bowtell, F.R. Balkwill, Nat. Rev. Cancer 11, 719–725 (2011). 10.1038/nrc3144 - PMC - PubMed
-
- M. Selvakumaran, D.A. Pisarcik, R. Bao, A.T. Yeung, T.C. Hamilton, Cancer Res. 63, 1311–1316 (2003) - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

