Dissection of tumoral niches using spatial transcriptomics and deep learning
- PMID: 40230519
- PMCID: PMC11994907
- DOI: 10.1016/j.isci.2025.112214
Dissection of tumoral niches using spatial transcriptomics and deep learning
Abstract
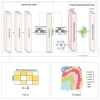
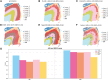
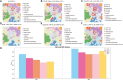
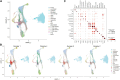
This study introduces TG-ME, an innovative computational framework that integrates transformer with graph variational autoencoder (GraphVAE) models for dissection of tumoral niches using spatial transcriptomics data and morphological images. TG-ME effectively identifies and characterizes niches in bench datasets and a high resolution NSCLC dataset. The pipeline consists in different stages that include normalization, spatial information integration, morphological feature extraction, gene expression quantification, single cell expression characterization, and tumor niche characterization. For this, TG-ME leverages advanced deep learning techniques that achieve robust clustering and profiling of niches across cancer stages. TG-ME can potentially provide insights into the spatial organization of tumor microenvironments (TME), highlighting specific niche compositions and their molecular changes along cancer progression. TG-ME is a promising tool for guiding personalized treatment strategies by uncovering microenvironmental signatures associated with disease prognosis and therapeutic outcomes.
Keywords: Artificial intelligence; Cancer; Microenvironment; Transcriptomics.
Conflict of interest statement
The authors declare no competing interests.
Figures






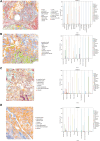
Similar articles
-
From tissue architecture to clinical insights: Spatial transcriptomics in solid tumor studies.Semin Oncol. 2025 Jul 14;52(5):152389. doi: 10.1016/j.seminoncol.2025.152389. Online ahead of print. Semin Oncol. 2025. PMID: 40664120 Review.
-
The Overlooked Role of Specimen Preparation in Bolstering Deep Learning-Enhanced Spatial Transcriptomics Workflows.medRxiv [Preprint]. 2023 Oct 9:2023.10.09.23296700. doi: 10.1101/2023.10.09.23296700. medRxiv. 2023. PMID: 37873287 Free PMC article. Preprint.
-
Gene spatial integration: enhancing spatial transcriptomics analysis via deep learning and batch effect mitigation.Bioinformatics. 2025 Jun 2;41(6):btaf350. doi: 10.1093/bioinformatics/btaf350. Bioinformatics. 2025. PMID: 40511994 Free PMC article.
-
Linking transcriptome and morphology in bone cells at cellular resolution with generative AI.J Bone Miner Res. 2024 Dec 31;40(1):20-26. doi: 10.1093/jbmr/zjae151. J Bone Miner Res. 2024. PMID: 39303095
-
A systematic review on feature extraction methods and deep learning models for detection of cancerous lung nodules at an early stage -the recent trends and challenges.Biomed Phys Eng Express. 2024 Nov 20;11(1). doi: 10.1088/2057-1976/ad9154. Biomed Phys Eng Express. 2024. PMID: 39530659
Cited by
-
Deciphering spatially confined immune evasion niches in osteosarcoma with 3-D spatial transcriptomics: a literature review.Front Oncol. 2025 Jul 16;15:1640645. doi: 10.3389/fonc.2025.1640645. eCollection 2025. Front Oncol. 2025. PMID: 40740859 Free PMC article. Review.
References
-
- Lorusso G., Rüegg C. The tumor microenvironment and its contribution to tumor evolution toward metastasis. Histochem. Cell Biol. 2008;130:1091–1103. - PubMed
-
- Kobayashi H., Enomoto A., Woods S.L., Burt A.D., Takahashi M., Worthley D.L. Cancer-associated fibroblasts in gastrointestinal cancer. Nat. Rev. Gastroenterol. Hepatol. 2019;16:282–295. - PubMed
-
- Mittal V., El Rayes T., Narula N., McGraw T.E., Altorki N.K., Barcellos-Hoff M.H. The microenvironment of lung cancer and therapeutic implications. Adv. Exp. Med. Biol. 2016;890:75–110. - PubMed
-
- Wang N., Li X., Wang R., Ding Z. Spatial transcriptomics and proteomics technologies for deconvoluting the tumor microenvironment. Biotechnol. J. 2021;16 - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

