Gut microbiome remodeling provides protection from an environmental toxin
- PMID: 40230520
- PMCID: PMC11995125
- DOI: 10.1016/j.isci.2025.112209
Gut microbiome remodeling provides protection from an environmental toxin
Abstract
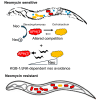
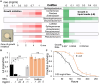
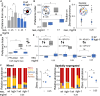
Gut microbiomes contribute to animal health and fitness. The immense biochemical diversity of bacteria holds particular potential for neutralizing environmental toxins and thus helping hosts deal with new toxic challenges. To explore this potential, we used Caenorhabditis elegans harboring a defined microbiome, and the antibiotic neomycin as a model toxin, differentially affecting microbiome strains, and also toxic to worms. Worms exposed to neomycin showed delayed development and reduced survival but were protected when colonized with neomycin-resistant Stenotrophomonas. 16S rRNA sequencing, bacterial load quantification, genetic manipulation, and behavioral assays showed that protection was linked to enrichment of Stenotrophomonas carrying a neomycin-modifying enzyme. Enrichment was facilitated by altered bacterial competition in the gut, as well as by KGB-1/JNK-dependent behavioral changes. While microbiome remodeling conferred toxin resistance, it was associated with reduced infection resistance and metabolic changes. These findings suggest that microbiome adaptation can help animals cope with stressors but may have long-term consequences that add to effects of direct intoxication.
Keywords: Microbiology; Microbiome.
© 2025 The Authors.
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Microbiome remodeling through bacterial competition and host behavior enables rapid adaptation to environmental toxins.bioRxiv [Preprint]. 2023 Jun 23:2023.06.21.545768. doi: 10.1101/2023.06.21.545768. bioRxiv. 2023. PMID: 37646003 Free PMC article. Preprint.
-
Identification and characterization of a skin microbiome on Caenorhabditis elegans suggests environmental microbes confer cuticle protection.Microbiol Spectr. 2024 Aug 6;12(8):e0016924. doi: 10.1128/spectrum.00169-24. Epub 2024 Jul 9. Microbiol Spectr. 2024. PMID: 38980017 Free PMC article.
-
Global landscape of gut microbiome diversity and antibiotic resistomes across vertebrates.Sci Total Environ. 2022 Sep 10;838(Pt 2):156178. doi: 10.1016/j.scitotenv.2022.156178. Epub 2022 May 23. Sci Total Environ. 2022. PMID: 35618126 Review.
-
Environmental Perturbations during the Rehabilitation of Wild Migratory Birds Induce Gut Microbiome Alteration and Antibiotic Resistance Acquisition.Microbiol Spectr. 2022 Aug 31;10(4):e0116322. doi: 10.1128/spectrum.01163-22. Epub 2022 Jun 22. Microbiol Spectr. 2022. PMID: 35730950 Free PMC article.
-
Microbiota and Autism: A Review on Oral and Gut Microbiome Analysis Through 16S rRNA Sequencing.Biomedicines. 2024 Nov 25;12(12):2686. doi: 10.3390/biomedicines12122686. Biomedicines. 2024. PMID: 39767593 Free PMC article. Review.
Cited by
-
Fecal microbiota transplantation in pigs: current status and future perspective.Anim Microbiome. 2025 Jul 20;7(1):76. doi: 10.1186/s42523-025-00440-w. Anim Microbiome. 2025. PMID: 40685369 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Research Materials

