Unveiling transcriptional mechanisms of B7-H3 in breast cancer stem cells through proteomic approaches
- PMID: 40230524
- PMCID: PMC11995042
- DOI: 10.1016/j.isci.2025.112218
Unveiling transcriptional mechanisms of B7-H3 in breast cancer stem cells through proteomic approaches
Abstract
B7-H3, an immune checkpoint molecule, is prominently overexpressed in various solid tumors, correlating with poor clinical outcomes. Despite its critical role in promoting tumorigenesis, metastasis, and immune evasion, the regulatory mechanisms governing B7-H3 expression, particularly in cancer stem cells (CSCs), remain elusive. In this comprehensive study, we focused on breast CSCs to uncover the transcriptional regulators driving B7-H3 overexpression. Utilizing DNA affinity purification-mass spectrometry (DAP-MS) to analyze B7-H3 promoter regions, we identified a novel set of transcription factors, including DDB1, XRCC5, PARP1, RPA1, and RPA3, as key modulators of B7-H3 expression. Functional assays revealed that targeting DDB1 with nitazoxanide significantly downregulated B7-H3 expression, subsequently impairing tumor sphere formation and cell migration in breast CSCs. These findings not only elucidate the complex transcriptional network controlling B7-H3 expression but also open new avenues for developing targeted immunotherapies aimed at disrupting CSC-driven cancer progression.
Keywords: cancer; cell biology; molecular biology; omics.
© 2025 The Author(s).
Conflict of interest statement
The authors declare no conflict of interest.
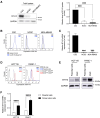
Figures






Similar articles
-
B7-H3 role in the immune landscape of cancer.Am J Clin Exp Immunol. 2017 Jun 15;6(4):66-75. eCollection 2017. Am J Clin Exp Immunol. 2017. PMID: 28695059 Free PMC article. Review.
-
To kill a cancer: Targeting the immune inhibitory checkpoint molecule, B7-H3.Biochim Biophys Acta Rev Cancer. 2022 Sep;1877(5):188783. doi: 10.1016/j.bbcan.2022.188783. Epub 2022 Aug 24. Biochim Biophys Acta Rev Cancer. 2022. PMID: 36028149 Review.
-
B7-H3 Expression in Breast Cancer and Brain Metastasis.Int J Mol Sci. 2024 Apr 3;25(7):3976. doi: 10.3390/ijms25073976. Int J Mol Sci. 2024. PMID: 38612786 Free PMC article.
-
Expression of B7-H3, a potential factor of tumor immune evasion in combination with the number of regulatory T cells, affects against recurrence-free survival in breast cancer patients.Ann Surg Oncol. 2014 Dec;21 Suppl 4(Suppl 4):S546-54. doi: 10.1245/s10434-014-3564-2. Epub 2014 Feb 22. Ann Surg Oncol. 2014. PMID: 24562936 Free PMC article.
-
Tumor Immunotherapy Targeting B7-H3: From Mechanisms to Clinical Applications.Immunotargets Ther. 2025 Mar 27;14:291-320. doi: 10.2147/ITT.S507522. eCollection 2025. Immunotargets Ther. 2025. PMID: 40171330 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

