Gut microbiome and short-chain fatty acids associated with the efficacy of growth hormone treatment in children with short stature
- PMID: 40230807
- PMCID: PMC11994682
- DOI: 10.3389/fped.2025.1557878
Gut microbiome and short-chain fatty acids associated with the efficacy of growth hormone treatment in children with short stature
Abstract
Objective: To investigate associations between fecal microbiota, short-chain fatty acids (SCFAs), and the efficacy of recombinant human growth hormone (rhGH) treatment in children with growth hormone deficiency (GHD) or idiopathic short stature (ISS).
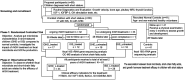
Methods: A 2-phase cohort study was conducted. Phase I included 102 participants (GHD: n = 33, ISS: n = 28, controls: n = 41) for cross-sectional analysis using 16S rRNA sequencing and targeted metabolomics to compare microbial diversity, predicted metabolic pathways, and SCFA levels. Phase II longitudinally monitored 61 rhGH-treated children (GHD = 33, ISS = 28) over 2 years, assessing growth velocity, IGF-1 levels, and fecal microbiota/SCFA dynamics. Statistical analyses included alpha/beta diversity metrics, LEfSe, PERMANOVA, and redundancy analysis (RDA) to link microbial/SCFA profiles with clinical outcomes.
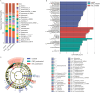
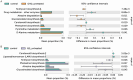
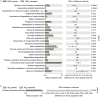
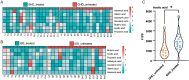
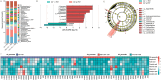
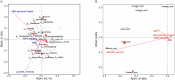
Results: (1). Microbiota Dysbiosis: Untreated GHD/ISS children exhibited reduced beneficial taxa (e.g., Faecalibacterium, Akkermansia) and increased pathobionts (e.g., Streptococcus, Collinsella) compared to controls (PERMANOVA: R 2 = 0.114, P = 0.001). (2). Metabolic Pathways: GHD/ISS groups showed enrichment in xenobiotic degradation (e.g., atrazine) and deficits in nutrient-associated pathways (e.g., carotenoid biosynthesis). (3). rhGH Effects: Treatment increased beneficial taxa (e.g., Bifidobacterium, Faecalibacterium) and modulated amino acid/lipid metabolism pathways (e.g., glycine-serine-threonine metabolism, P = 0.035). (4). SCFAs and Growth Velocity: Higher growth velocity percentiles correlated with elevated acetic acid (GHD-treated: 1952 ± 962.4 vs. untreated: 1290 ± 886.0 μg/g, P = 0.037) and butyric acid levels.
Conclusion: GHD, ISS, and healthy children have different fecal microbiota compositions and SCFA metabolisms. rhGH therapy partially restores microbial balance and alters metabolic pathways, with SCFA levels associated with treatment efficacy. These findings highlight the gut microbiome as a potential modulator of rhGH response and provide insight into microbiota-targeted therapies to improve growth outcomes (e.g., "probiotic interventions").
Keywords: IGF-1 (insulin-like growth factor 1); SCFAs (short-chain fatty acids); growth hormone treatment; microbiota; short stature.
© 2025 He, Lyu, Shen, Liu, Zhang, Li, Huang, Xu, Zhang and Guo.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial ties that might be perceived as a potential conflict of interest.
Figures


Similar articles
-
Impact of different growth hormone levels on gut microbiota and metabolism in short stature.Pediatr Res. 2024 Jul;96(1):115-123. doi: 10.1038/s41390-024-03140-4. Epub 2024 Apr 6. Pediatr Res. 2024. PMID: 38582946
-
A Real-World Study of Recombinant Human Growth Hormone in the Treatment of Idiopathic Short Stature and Growth Hormone Deficiency.Ther Clin Risk Manag. 2022 Mar 16;18:113-124. doi: 10.2147/TCRM.S363564. eCollection 2022. Ther Clin Risk Manag. 2022. PMID: 35342293 Free PMC article.
-
Characteristics of Gut Microbiome and Its Metabolites, Short-Chain Fatty Acids, in Children With Idiopathic Short Stature.Front Endocrinol (Lausanne). 2022 Jun 10;13:890200. doi: 10.3389/fendo.2022.890200. eCollection 2022. Front Endocrinol (Lausanne). 2022. PMID: 35757432 Free PMC article.
-
Comparison of the efficacy and safety of recombinant human growth hormone in treating idiopathic short stature and growth hormone deficiency in children.Growth Horm IGF Res. 2020 Aug-Oct;53-54:101331. doi: 10.1016/j.ghir.2020.101331. Epub 2020 Jul 17. Growth Horm IGF Res. 2020. PMID: 32777706
-
Gut-brain-liver axis in growth hormone deficiency: role of microbiota-derived short-chain fatty acids in ethnic variability and therapeutic development.Front Public Health. 2025 May 30;13:1541654. doi: 10.3389/fpubh.2025.1541654. eCollection 2025. Front Public Health. 2025. PMID: 40520290 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

