A circRNA-mRNA pairing mechanism regulates tumor growth and endocrine therapy resistance in ER-positive breast cancer
- PMID: 40233410
- PMCID: PMC11874584
- DOI: 10.1073/pnas.2420383122
A circRNA-mRNA pairing mechanism regulates tumor growth and endocrine therapy resistance in ER-positive breast cancer
Abstract
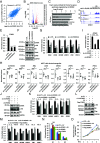
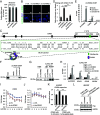
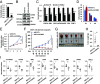
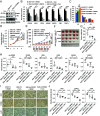
The molecular mechanisms underlying estrogen receptor (ER)-positive breast carcinogenesis and drug resistance remain incompletely understood. Elevated expression of CCND1 is linked to enhanced invasiveness, poorer prognosis, and resistance to drug therapies in ER-positive breast cancer. In this study, we identify a highly expressed circular RNA (circRNA) derived from FOXK2, called circFOXK2, which plays a key role in stabilizing CCND1 mRNA, thereby promoting cell cycle progression, cell growth, and endocrine therapy resistance in ER-positive breast cancer cells. Mechanistically, circFOXK2 binds directly to CCND1 mRNA via RNA-RNA pairing and recruits the RNA-binding protein ELAVL1/HuR, stabilizing the CCND1 mRNA and enhancing CCND1 protein levels. This results in activation of the CCND1-CDK4/6-p-RB-E2F signaling axis, driving the transcription of downstream E2F target genes and facilitating the G1/S transition during cell cycle progression. Notably, targeting circFOXK2 with antisense oligonucleotide (ASO-circFOXK2) suppresses ER-positive breast cancer cell growth both in vitro and in vivo. Moreover, combination therapy with ASO-circFOXK2 and tamoxifen exhibits synergistic effects and restores tamoxifen sensitivity in tamoxifen-resistant cells. Clinically, high circFOXK2 expression is positively correlated with CCND1 levels in both ER-positive breast cancer cell lines and patient tumor tissues. Overall, our findings reveal the critical role of circFOXK2 in stabilizing the oncogene CCND1 and promoting cancer progression, positioning circFOXK2 as a potential therapeutic target for ER-positive breast cancer in clinical settings.
Keywords: CCND1; RNA-RNA pairing; breast cancer; circFOXK2; endocrine therapy resistance.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures


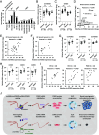
Similar articles
-
Post-transcriptional regulation of ERBB2 by miR26a/b and HuR confers resistance to tamoxifen in estrogen receptor-positive breast cancer cells.J Biol Chem. 2017 Aug 18;292(33):13551-13564. doi: 10.1074/jbc.M117.780973. Epub 2017 Jun 21. J Biol Chem. 2017. PMID: 28637868 Free PMC article.
-
MICAL-L2, as an estrogen-responsive gene, is involved in ER-positive breast cancer cell progression and tamoxifen sensitivity via the AKT/mTOR pathway.Biochem Pharmacol. 2024 Jul;225:116256. doi: 10.1016/j.bcp.2024.116256. Epub 2024 May 9. Biochem Pharmacol. 2024. PMID: 38729448
-
Estrogen-induced circRNA, circPGR, functions as a ceRNA to promote estrogen receptor-positive breast cancer cell growth by regulating cell cycle-related genes.Theranostics. 2021 Jan 1;11(4):1732-1752. doi: 10.7150/thno.45302. eCollection 2021. Theranostics. 2021. PMID: 33408778 Free PMC article.
-
circRNA circ-CCND1 promotes the proliferation of laryngeal squamous cell carcinoma through elevating CCND1 expression via interacting with HuR and miR-646.J Cell Mol Med. 2020 Feb;24(4):2423-2433. doi: 10.1111/jcmm.14925. Epub 2020 Jan 17. J Cell Mol Med. 2020. PMID: 31951319 Free PMC article.
-
99mTc-CCND1-PNA-IGF1 Chimeras.2006 Nov 1 [updated 2008 Jan 16]. In: Molecular Imaging and Contrast Agent Database (MICAD) [Internet]. Bethesda (MD): National Center for Biotechnology Information (US); 2004–2013. 2006 Nov 1 [updated 2008 Jan 16]. In: Molecular Imaging and Contrast Agent Database (MICAD) [Internet]. Bethesda (MD): National Center for Biotechnology Information (US); 2004–2013. PMID: 20641356 Free Books & Documents. Review.
Cited by
-
Circular RNAs as Targets for Developing Anticancer Therapeutics.Cells. 2025 Jul 18;14(14):1106. doi: 10.3390/cells14141106. Cells. 2025. PMID: 40710359 Free PMC article. Review.
-
Direct circRNA-mRNA Binding Controls mRNA Fate: A New Mechanism for circRNAs.Noncoding RNA. 2025 Jul 18;11(4):53. doi: 10.3390/ncrna11040053. Noncoding RNA. 2025. PMID: 40700095 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
- 82125028/MOST | National Natural Science Foundation of China (NSFC)
- U22A20320/MOST | National Natural Science Foundation of China (NSFC)
- 31871319/MOST | National Natural Science Foundation of China (NSFC)
- 91953114/MOST | National Natural Science Foundation of China (NSFC)
- 2020YFA0112300/Ministry of Science and Technology of the People's Republic of China (MOST)
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

